cTRAP: identifying candidate causal perturbations from differential gene expression data
Bernardo P. de Almeida & Nuno Saraiva-Agostinho
2026-01-12
Source:vignettes/cTRAP.Rmd
cTRAP.RmdIntroduction
cTRAP is an R package designed to compare differential
gene expression results with those from known cellular perturbations
(such as gene knockdown, overexpression or small molecules) derived from
the Connectivity Map (CMap; Subramanian et al.,
Cell 2017). Such analyses allow not only to infer the molecular
causes of the observed difference in gene expression but also to
identify small molecules that could drive or revert specific
transcriptomic alterations.
To illustrate the package functionalities, we will use an example based on a gene knockdown dataset from the ENCODE project for which there is available RNA-seq data. After performing differential expression analyses to the matched-control sample, we will compare the respective transcriptomic changes with the ones caused by all CMap’s gene knockdown perturbations to identify which ones have similar or inverse transcriptomic changes to the observed ones. As a positive control, we expect to find the knock-down of the gene depleted in the ENCODE experiment as one of the most similar transcriptomic perturbations.
Getting started
To load the cTRAP package into your R environment
type:
## Registered S3 method overwritten by 'quantmod':
## method from
## as.zoo.data.frame zooLoad ENCODE RNA-seq data and perform differential gene expression analysis
In this example, we will use the EIF4G1 shRNA knockdown followed by RNA-seq experiment in HepG2 cell line from the ENCODE project as the dataset of interest. The RNA-seq processed data (gene quantifications from RSEM method) for the EIF4G1 knock-down and respective controls (two replicates each) can be automatically downloaded and loaded by typing:
gene <- "EIF4G1"
cellLine <- "HepG2"
ENCODEmetadata <- downloadENCODEknockdownMetadata(cellLine, gene)
table(ENCODEmetadata$`Experiment target`)
length(unique(ENCODEmetadata$`Experiment target`))
ENCODEsamples <- loadENCODEsamples(ENCODEmetadata)[[1]]
counts <- prepareENCODEgeneExpression(ENCODEsamples)Gene expression data (read counts) were quantile-normalized using voom
and differential expression analysis was performed using the limma
R package.
# Remove low coverage (at least 10 counts shared across two samples)
minReads <- 10
minSamples <- 2
filter <- rowSums(counts[ , -c(1, 2)] >= minReads) >= minSamples
counts <- counts[filter, ]
# Convert ENSEMBL identifier to gene symbol
counts$gene_id <- convertGeneIdentifiers(counts$gene_id)
# Perform differential gene expression analysis
diffExpr <- performDifferentialExpression(counts)For our metric of differential expression after EIF4G1 shRNA knock-down, we will use the respective t-statistic.
# Get t-statistics of differential expression with respective gene names
# (expected input for downstream analyses)
diffExprStat <- diffExpr$t
names(diffExprStat) <- diffExpr$Gene_symbolLoad CMap perturbation data
We will use our differential gene expression metric to compare with CMap’s gene knock-down perturbations in the same cell line (HepG2). Note that this comparison can also be done to perturbations in a different cell line (or in all cell lines using the average result across cell lines).
To summarise conditions and check available data in CMap, we can use the following commands to download CMap metadata:
# Load CMap metadata (automatically downloaded if not found)
cmapMetadata <- loadCMapData("cmapMetadata.txt", type="metadata")## cmapMetadata.txt not found: downloading data...## Extracting cmapMetadata.txt.gz...## Loading CMap metadata from cmapMetadata.txt...
# Summarise conditions for all CMap perturbations
getCMapConditions(cmapMetadata)## $perturbationType
## [1] "Compound"
## [2] "Peptides and other biological agents (e.g. cytokine)"
## [3] "shRNA for loss of function (LoF) of gene"
## [4] "Consensus signature from shRNAs targeting the same gene"
## [5] "cDNA for overexpression of wild-type gene"
## [6] "cDNA for overexpression of mutated gene"
##
## $cellLine
## [1] "A375" "A549" "A673" "AGS" "ASC" "BT20"
## [7] "CD34" "CL34" "CORL23" "COV644" "DV90" "EFO27"
## [13] "FIBRNPC" "H1299" "HA1E" "HCC15" "HCC515" "HCT116"
## [19] "HEC108" "HEK293T" "HEKTE" "HEPG2" "HL60" "HS27A"
## [25] "HS578T" "HT115" "HT29" "HUH7" "JHUEM2" "JURKAT"
## [31] "LOVO" "MCF10A" "MCF7" "MCH58" "MDAMB231" "MDST8"
## [37] "NCIH1694" "NCIH1836" "NCIH2073" "NCIH508" "NCIH596" "NCIH716"
## [43] "NEU" "NKDBA" "NOMO1" "NPC" "OV7" "PC3"
## [49] "PHH" "PL21" "RKO" "RMGI" "RMUGS" "SHSY5Y"
## [55] "SKB" "SKBR3" "SKL" "SKLU1" "SKM1" "SKMEL1"
## [61] "SKMEL28" "SNGM" "SNU1040" "SNUC4" "SNUC5" "SW480"
## [67] "SW620" "SW948" "T3M10" "THP1" "TYKNU" "U266"
## [73] "U2OS" "U937" "VCAP" "WSUDLCL2"
##
## $dosage
## [1] NA "0.1 %" "1 nM" "10 nM" "100 nM"
## [6] "500 nM" "0.04 µM" "0.12 µM" "0.37 µM" "0.41 µM"
## [11] "1 µM" "1.11 µM" "1.23 µM" "3 µM" "3.33 µM"
## [16] "3.7 µM" "5 µM" "10 µM" "11.11 µM" "20 µM"
## [21] "30 µM" "33.33 µM" "40 µM" "50 µM" "60 µM"
## [26] "70 µM" "80 µM" "90 µM" "100 µM" "1 µL"
## [31] "1.5 µL" "2 µL" "3 µL" "4 µL" "5 µL"
## [36] "6 µL" "10 µL" "20 µL" "150 ng" "200 ng"
## [41] "300 ng" "0.1 ng/µL" "1 ng/µL" "3 ng/µL" "10 ng/µL"
## [46] "100 ng/µL" "300 ng/µL" "0 ng/mL" "0.01 ng/mL" "0.03 ng/mL"
## [51] "0.1 ng/mL" "0.15 ng/mL" "0.2 ng/mL" "0.3 ng/mL" "0.5 ng/mL"
## [56] "1 ng/mL" "1.65 ng/mL" "2 ng/mL" "5 ng/mL" "10 ng/mL"
## [61] "15 ng/mL" "16 ng/mL" "20 ng/mL" "25 ng/mL" "30 ng/mL"
## [66] "40 ng/mL" "45 ng/mL" "50 ng/mL" "80 ng/mL" "100 ng/mL"
## [71] "150 ng/mL" "200 ng/mL" "250 ng/mL" "400 ng/mL" "500 ng/mL"
## [76] "800 ng/mL" "1000 ng/mL" "2000 ng/mL" "2500 ng/mL" "3000 ng/mL"
## [81] "5000 ng/mL" "8300 ng/mL" "10000 ng/mL" "50000 ng/mL" "100000 ng/mL"
## [86] "200000 ng/mL"
##
## $timepoint
## [1] "1 h" "2 h" "3 h" "4 h" "6 h" "24 h" "48 h" "72 h" "96 h"
## [10] "120 h" "144 h" "168 h"
# Summarise conditions for CMap perturbations in HepG2 cell line
getCMapConditions(cmapMetadata, cellLine="HepG2")## $perturbationType
## [1] "Compound"
## [2] "Peptides and other biological agents (e.g. cytokine)"
## [3] "shRNA for loss of function (LoF) of gene"
## [4] "Consensus signature from shRNAs targeting the same gene"
## [5] "cDNA for overexpression of wild-type gene"
##
## $cellLine
## [1] "HEPG2"
##
## $dosage
## [1] NA "0.1 %" "10 nM" "100 nM" "500 nM"
## [6] "0.04 µM" "0.12 µM" "0.37 µM" "0.41 µM" "1 µM"
## [11] "1.11 µM" "1.23 µM" "3 µM" "3.33 µM" "3.7 µM"
## [16] "5 µM" "10 µM" "11.11 µM" "20 µM" "30 µM"
## [21] "33.33 µM" "40 µM" "50 µM" "60 µM" "70 µM"
## [26] "80 µM" "90 µM" "100 µM" "1.5 µL" "2 µL"
## [31] "0 ng/mL" "0.01 ng/mL" "0.03 ng/mL" "0.1 ng/mL" "0.15 ng/mL"
## [36] "0.2 ng/mL" "0.3 ng/mL" "0.5 ng/mL" "1 ng/mL" "1.65 ng/mL"
## [41] "2 ng/mL" "5 ng/mL" "10 ng/mL" "15 ng/mL" "16 ng/mL"
## [46] "20 ng/mL" "25 ng/mL" "30 ng/mL" "40 ng/mL" "45 ng/mL"
## [51] "50 ng/mL" "80 ng/mL" "100 ng/mL" "150 ng/mL" "200 ng/mL"
## [56] "250 ng/mL" "400 ng/mL" "500 ng/mL" "800 ng/mL" "1000 ng/mL"
## [61] "2000 ng/mL" "2500 ng/mL" "3000 ng/mL" "5000 ng/mL" "8300 ng/mL"
## [66] "10000 ng/mL" "50000 ng/mL" "100000 ng/mL" "200000 ng/mL"
##
## $timepoint
## [1] "2 h" "4 h" "6 h" "24 h" "96 h"
# Summarise conditions for a specific CMap perturbation in HepG2 cell line
getCMapConditions(
cmapMetadata, cellLine="HepG2",
perturbationType="Consensus signature from shRNAs targeting the same gene")## $perturbationType
## [1] "Consensus signature from shRNAs targeting the same gene"
##
## $cellLine
## [1] "HEPG2"
##
## $dosage
## [1] "1.5 µL"
##
## $timepoint
## [1] "96 h"Now, we will filter the metadata to CMap gene knockdown perturbations in HepG2 and load associated gene information and differential gene expression data based on the given filename (the file is automatically downloaded if it does not exist).
Differential gene expression data for each CMap perturbation are available in normalised z-score values (read Subramanian et al., Cell 2017 for more details). As the file is big (around 20GB), a prompt will ask to confirm whether to download the file. If you prefer, you can also download the file yourself:
- Manually download the file from GSE92742_Broad_LINCS_Level5_COMPZ.MODZ_n473647x12328.gctx.gz
- Verify file integrity after download using the SHA-512 checksum
- Use the exact same filename for the
zscoresargument ofprepareCMapPerturbations())
# Filter CMap gene knockdown HepG2 data to be loaded
cmapMetadataKD <- filterCMapMetadata(
cmapMetadata, cellLine="HepG2",
perturbationType="Consensus signature from shRNAs targeting the same gene")
# Load filtered data (data will be downloaded if not found)
cmapPerturbationsKD <- prepareCMapPerturbations(
metadata=cmapMetadataKD, zscores="cmapZscores.gctx",
geneInfo="cmapGeneInfo.txt")If interested in small molecules, the differential gene expression z-scores from CMap can be downloaded for each small molecule perturbation:
# Filter CMap gene small molecule HepG2 data to be loaded
cmapMetadataCompounds <- filterCMapMetadata(
cmapMetadata, cellLine="HepG2", perturbationType="Compound")
# Load filtered data (data will be downloaded if not found)
cmapPerturbationsCompounds <- prepareCMapPerturbations(
metadata=cmapMetadataCompounds, zscores="cmapZscores.gctx",
geneInfo="cmapGeneInfo.txt", compoundInfo="cmapCompoundInfo.txt")Comparison with CMap perturbations
The rankSimilarPerturbations function compares the
differential expression metric (the t-statistic, in this case) against
the CMap perturbations’ z-scores using the available methods:
- Spearman’s correlation
- Pearson’s correlation
- Gene Set Enrichment Analysis (GSEA), where the most up- and down-regulated n genes from the user’s differential expression profile are used as gene sets (by default, n = 150 genes)
To compare against CMap knockdown perturbations using all the previous methods:
compareKD <- rankSimilarPerturbations(diffExprStat, cmapPerturbationsKD)To compare against selected CMap small molecule perturbations:
compareCompounds <- rankSimilarPerturbations(diffExprStat,
cmapPerturbationsCompounds)The output table contains the results of the comparison with each perturbation tested, including the test statistics (Spearman’s correlation coefficient, Pearson’s correlation coefficient and/or GSEA score), the respective p-value and the Benjamini-Hochberg-adjusted p-value (for correlation statistics only). When performing multiple methods, the rank product’s rank will be included to summarise other method’s rankings.
# Most positively associated perturbations (note that EIF4G1 knockdown is the
# 7th, 1st and 2nd most positively associated perturbation based on Spearman's
# correlation, Pearson's correlation and GSEA, respectively)
head(compareKD[order(spearman_rank)], n=10)## gene_perturbation spearman_coef spearman_pvalue spearman_qvalue
## <char> <num> <num> <num>
## 1: CGS001_HEPG2_96H:MECP2:1.5 0.1930335 1.506067e-74 1.305258e-73
## 2: CGS001_HEPG2_96H:KIAA0196:1.5 0.1894301 8.237226e-72 5.354197e-71
## 3: CGS001_HEPG2_96H:SQRDL:1.5 0.1883210 5.589838e-71 2.906716e-70
## 4: CGS001_HEPG2_96H:PPIH:1.5 0.1863954 1.509735e-69 6.542184e-69
## 5: CGS001_HEPG2_96H:STAT1:1.5 0.1833382 2.626628e-67 9.756048e-67
## 6: CGS001_HEPG2_96H:COPS5:1.5 0.1775880 3.364592e-63 1.093493e-62
## 7: CGS001_HEPG2_96H:EIF4G1:1.5 0.1770993 7.406639e-63 2.139696e-62
## 8: CGS001_HEPG2_96H:MEST:1.5 0.1726479 8.816544e-60 2.292301e-59
## 9: CGS001_HEPG2_96H:KIF20A:1.5 0.1665159 1.114839e-55 2.415485e-55
## 10: CGS001_HEPG2_96H:TMEM5:1.5 0.1506335 8.936446e-46 1.452172e-45
## pearson_coef pearson_pvalue pearson_qvalue GSEA spearman_rank
## <num> <num> <num> <num> <num>
## 1: 0.1756788 7.245930e-62 1.883942e-61 0.3903862 1
## 2: 0.1847352 2.514633e-68 9.340065e-68 0.3521034 2
## 3: 0.1718559 3.046859e-59 6.093718e-59 0.3998024 3
## 4: 0.1703630 3.104268e-58 5.380731e-58 0.4359134 4
## 5: 0.1894661 7.739198e-72 4.024383e-71 0.3765351 5
## 6: 0.1794790 1.553956e-64 5.050356e-64 0.4222756 6
## 7: 0.1915142 2.182595e-73 1.418686e-72 0.4861485 7
## 8: 0.1858475 3.830708e-69 1.659973e-68 0.4219693 8
## 9: 0.1786967 5.568601e-64 1.608707e-63 0.4566918 9
## 10: 0.1670226 5.177976e-56 8.414211e-56 0.4721782 10
## pearson_rank GSEA_rank rankProduct_rank
## <num> <num> <num>
## 1: 7 12 2
## 2: 4 14 4
## 3: 9 11 8
## 4: 10 8 9
## 5: 2 13 5
## 6: 5 9 7
## 7: 1 2 1
## 8: 3 10 6
## 9: 6 7 10
## 10: 11 6 12## gene_perturbation spearman_coef spearman_pvalue spearman_qvalue
## <char> <num> <num> <num>
## 1: CGS001_HEPG2_96H:EIF4G1:1.5 0.1770993 7.406639e-63 2.139696e-62
## 2: CGS001_HEPG2_96H:STAT1:1.5 0.1833382 2.626628e-67 9.756048e-67
## 3: CGS001_HEPG2_96H:MEST:1.5 0.1726479 8.816544e-60 2.292301e-59
## 4: CGS001_HEPG2_96H:KIAA0196:1.5 0.1894301 8.237226e-72 5.354197e-71
## 5: CGS001_HEPG2_96H:COPS5:1.5 0.1775880 3.364592e-63 1.093493e-62
## 6: CGS001_HEPG2_96H:KIF20A:1.5 0.1665159 1.114839e-55 2.415485e-55
## pearson_coef pearson_pvalue pearson_qvalue GSEA spearman_rank
## <num> <num> <num> <num> <num>
## 1: 0.1915142 2.182595e-73 1.418686e-72 0.4861485 7
## 2: 0.1894661 7.739198e-72 4.024383e-71 0.3765351 5
## 3: 0.1858475 3.830708e-69 1.659973e-68 0.4219693 8
## 4: 0.1847352 2.514633e-68 9.340065e-68 0.3521034 2
## 5: 0.1794790 1.553956e-64 5.050356e-64 0.4222756 6
## 6: 0.1786967 5.568601e-64 1.608707e-63 0.4566918 9
## pearson_rank GSEA_rank rankProduct_rank
## <num> <num> <num>
## 1: 1 2 1
## 2: 2 13 5
## 3: 3 10 6
## 4: 4 14 4
## 5: 5 9 7
## 6: 6 7 10## gene_perturbation spearman_coef spearman_pvalue spearman_qvalue
## <char> <num> <num> <num>
## 1: CGS001_HEPG2_96H:SKIV2L:1.5 0.1423207 5.305914e-41 8.114927e-41
## 2: CGS001_HEPG2_96H:EIF4G1:1.5 0.1770993 7.406639e-63 2.139696e-62
## 3: CGS001_HEPG2_96H:GTPBP8:1.5 0.1190220 4.210973e-29 4.760230e-29
## 4: CGS001_HEPG2_96H:HFE:1.5 0.1235293 3.098180e-31 3.661486e-31
## 5: CGS001_HEPG2_96H:UBAP2L:1.5 0.1309546 6.320554e-35 8.216721e-35
## 6: CGS001_HEPG2_96H:TMEM5:1.5 0.1506335 8.936446e-46 1.452172e-45
## pearson_coef pearson_pvalue pearson_qvalue GSEA spearman_rank
## <num> <num> <num> <num> <num>
## 1: 0.1755377 9.078490e-62 2.145825e-61 0.5070024 11
## 2: 0.1915142 2.182595e-73 1.418686e-72 0.4861485 7
## 3: 0.1390818 3.229116e-39 3.816228e-39 0.4813972 14
## 4: 0.1499527 2.252417e-45 2.788707e-45 0.4813046 13
## 5: 0.1517482 1.948821e-46 2.533468e-46 0.4722054 12
## 6: 0.1670226 5.177976e-56 8.414211e-56 0.4721782 10
## pearson_rank GSEA_rank rankProduct_rank
## <num> <num> <num>
## 1: 8 1 3
## 2: 1 2 1
## 3: 14 3 11
## 4: 13 4 13
## 5: 12 5 14
## 6: 11 6 12## gene_perturbation spearman_coef spearman_pvalue spearman_qvalue
## <char> <num> <num> <num>
## 1: CGS001_HEPG2_96H:EIF4G1:1.5 0.1770993 7.406639e-63 2.139696e-62
## 2: CGS001_HEPG2_96H:MECP2:1.5 0.1930335 1.506067e-74 1.305258e-73
## 3: CGS001_HEPG2_96H:SKIV2L:1.5 0.1423207 5.305914e-41 8.114927e-41
## 4: CGS001_HEPG2_96H:KIAA0196:1.5 0.1894301 8.237226e-72 5.354197e-71
## 5: CGS001_HEPG2_96H:STAT1:1.5 0.1833382 2.626628e-67 9.756048e-67
## 6: CGS001_HEPG2_96H:MEST:1.5 0.1726479 8.816544e-60 2.292301e-59
## pearson_coef pearson_pvalue pearson_qvalue GSEA spearman_rank
## <num> <num> <num> <num> <num>
## 1: 0.1915142 2.182595e-73 1.418686e-72 0.4861485 7
## 2: 0.1756788 7.245930e-62 1.883942e-61 0.3903862 1
## 3: 0.1755377 9.078490e-62 2.145825e-61 0.5070024 11
## 4: 0.1847352 2.514633e-68 9.340065e-68 0.3521034 2
## 5: 0.1894661 7.739198e-72 4.024383e-71 0.3765351 5
## 6: 0.1858475 3.830708e-69 1.659973e-68 0.4219693 8
## pearson_rank GSEA_rank rankProduct_rank
## <num> <num> <num>
## 1: 1 2 1
## 2: 7 12 2
## 3: 8 1 3
## 4: 4 14 4
## 5: 2 13 5
## 6: 3 10 6## gene_perturbation spearman_coef spearman_pvalue spearman_qvalue
## <char> <num> <num> <num>
## 1: CGS001_HEPG2_96H:EYA1:1.5 -0.1993531 1.747554e-79 4.543641e-78
## 2: CGS001_HEPG2_96H:CDCA8:1.5 -0.1941862 1.951544e-75 2.537007e-74
## 3: CGS001_HEPG2_96H:NDUFB6:1.5 -0.1677880 1.618399e-56 3.825307e-56
## 4: CGS001_HEPG2_96H:EHMT2:1.5 -0.1640525 4.479303e-54 8.958605e-54
## 5: CGS001_HEPG2_96H:ZBTB24:1.5 -0.1632375 1.500986e-53 2.787546e-53
## 6: CGS001_HEPG2_96H:MAF:1.5 -0.1581847 2.351511e-50 4.075952e-50
## pearson_coef pearson_pvalue pearson_qvalue GSEA spearman_rank
## <num> <num> <num> <num> <num>
## 1: -0.1952303 3.030978e-76 7.880542e-75 -0.3776731 26
## 2: -0.1926944 2.740646e-74 2.375227e-73 -0.4739211 25
## 3: -0.1646057 1.964288e-54 3.004205e-54 0.0000000 24
## 4: -0.1590654 6.637816e-51 9.587957e-51 -0.3716818 23
## 5: -0.1948077 6.448588e-76 8.383165e-75 0.0000000 22
## 6: -0.1717844 3.406704e-59 6.326737e-59 0.0000000 21
## pearson_rank GSEA_rank rankProduct_rank
## <num> <num> <num>
## 1: 26 19 25
## 2: 24 26 26
## 3: 21 16 20
## 4: 20 18 21
## 5: 25 16 22
## 6: 22 16 19## gene_perturbation spearman_coef spearman_pvalue spearman_qvalue
## <char> <num> <num> <num>
## 1: CGS001_HEPG2_96H:EYA1:1.5 -0.1993531 1.747554e-79 4.543641e-78
## 2: CGS001_HEPG2_96H:ZBTB24:1.5 -0.1632375 1.500986e-53 2.787546e-53
## 3: CGS001_HEPG2_96H:CDCA8:1.5 -0.1941862 1.951544e-75 2.537007e-74
## 4: CGS001_HEPG2_96H:SULT1A2:1.5 -0.1414914 1.533416e-40 2.214934e-40
## 5: CGS001_HEPG2_96H:MAF:1.5 -0.1581847 2.351511e-50 4.075952e-50
## 6: CGS001_HEPG2_96H:NDUFB6:1.5 -0.1677880 1.618399e-56 3.825307e-56
## pearson_coef pearson_pvalue pearson_qvalue GSEA spearman_rank
## <num> <num> <num> <num> <num>
## 1: -0.1952303 3.030978e-76 7.880542e-75 -0.3776731 26
## 2: -0.1948077 6.448588e-76 8.383165e-75 0.0000000 22
## 3: -0.1926944 2.740646e-74 2.375227e-73 -0.4739211 25
## 4: -0.1742848 6.666242e-61 1.444352e-60 -0.4086520 20
## 5: -0.1717844 3.406704e-59 6.326737e-59 0.0000000 21
## 6: -0.1646057 1.964288e-54 3.004205e-54 0.0000000 24
## pearson_rank GSEA_rank rankProduct_rank
## <num> <num> <num>
## 1: 26 19 25
## 2: 25 16 22
## 3: 24 26 26
## 4: 23 20 24
## 5: 22 16 19
## 6: 21 16 20## gene_perturbation spearman_coef spearman_pvalue spearman_qvalue
## <char> <num> <num> <num>
## 1: CGS001_HEPG2_96H:CDCA8:1.5 -0.19418619 1.951544e-75 2.537007e-74
## 2: CGS001_HEPG2_96H:DHX16:1.5 -0.14109904 2.527811e-40 3.459110e-40
## 3: CGS001_HEPG2_96H:SIAH2:1.5 -0.07825242 2.030400e-13 2.111616e-13
## 4: CGS001_HEPG2_96H:GNAI2:1.5 -0.05956052 2.293513e-08 2.293513e-08
## 5: CGS001_HEPG2_96H:SHB:1.5 -0.11094246 1.768452e-25 1.915823e-25
## 6: CGS001_HEPG2_96H:PLOD2:1.5 -0.12393601 1.970762e-31 2.439991e-31
## pearson_coef pearson_pvalue pearson_qvalue GSEA spearman_rank
## <num> <num> <num> <num> <num>
## 1: -0.19269443 2.740646e-74 2.375227e-73 -0.4739211 25
## 2: -0.15642416 2.883756e-49 3.946193e-49 -0.4379221 19
## 3: -0.11367900 1.119976e-26 1.164775e-26 -0.4224346 16
## 4: -0.08923031 5.233713e-17 5.233713e-17 -0.4166373 15
## 5: -0.12875045 8.300805e-34 8.992538e-34 -0.4131576 17
## 6: -0.13259151 9.070613e-36 1.025374e-35 -0.4130739 18
## pearson_rank GSEA_rank rankProduct_rank
## <num> <num> <num>
## 1: 24 26 26
## 2: 19 25 23
## 3: 16 24 16
## 4: 15 23 15
## 5: 17 22 17
## 6: 18 21 18## gene_perturbation spearman_coef spearman_pvalue spearman_qvalue
## <char> <num> <num> <num>
## 1: CGS001_HEPG2_96H:CDCA8:1.5 -0.1941862 1.951544e-75 2.537007e-74
## 2: CGS001_HEPG2_96H:EYA1:1.5 -0.1993531 1.747554e-79 4.543641e-78
## 3: CGS001_HEPG2_96H:SULT1A2:1.5 -0.1414914 1.533416e-40 2.214934e-40
## 4: CGS001_HEPG2_96H:DHX16:1.5 -0.1410990 2.527811e-40 3.459110e-40
## 5: CGS001_HEPG2_96H:ZBTB24:1.5 -0.1632375 1.500986e-53 2.787546e-53
## 6: CGS001_HEPG2_96H:EHMT2:1.5 -0.1640525 4.479303e-54 8.958605e-54
## pearson_coef pearson_pvalue pearson_qvalue GSEA spearman_rank
## <num> <num> <num> <num> <num>
## 1: -0.1926944 2.740646e-74 2.375227e-73 -0.4739211 25
## 2: -0.1952303 3.030978e-76 7.880542e-75 -0.3776731 26
## 3: -0.1742848 6.666242e-61 1.444352e-60 -0.4086520 20
## 4: -0.1564242 2.883756e-49 3.946193e-49 -0.4379221 19
## 5: -0.1948077 6.448588e-76 8.383165e-75 0.0000000 22
## 6: -0.1590654 6.637816e-51 9.587957e-51 -0.3716818 23
## pearson_rank GSEA_rank rankProduct_rank
## <num> <num> <num>
## 1: 24 26 26
## 2: 26 19 25
## 3: 23 20 24
## 4: 19 25 23
## 5: 25 16 22
## 6: 20 18 21## Warning: No shared levels found between `names(values)` of the manual scale and the
## data's colour values.
## No shared levels found between `names(values)` of the manual scale and the
## data's colour values.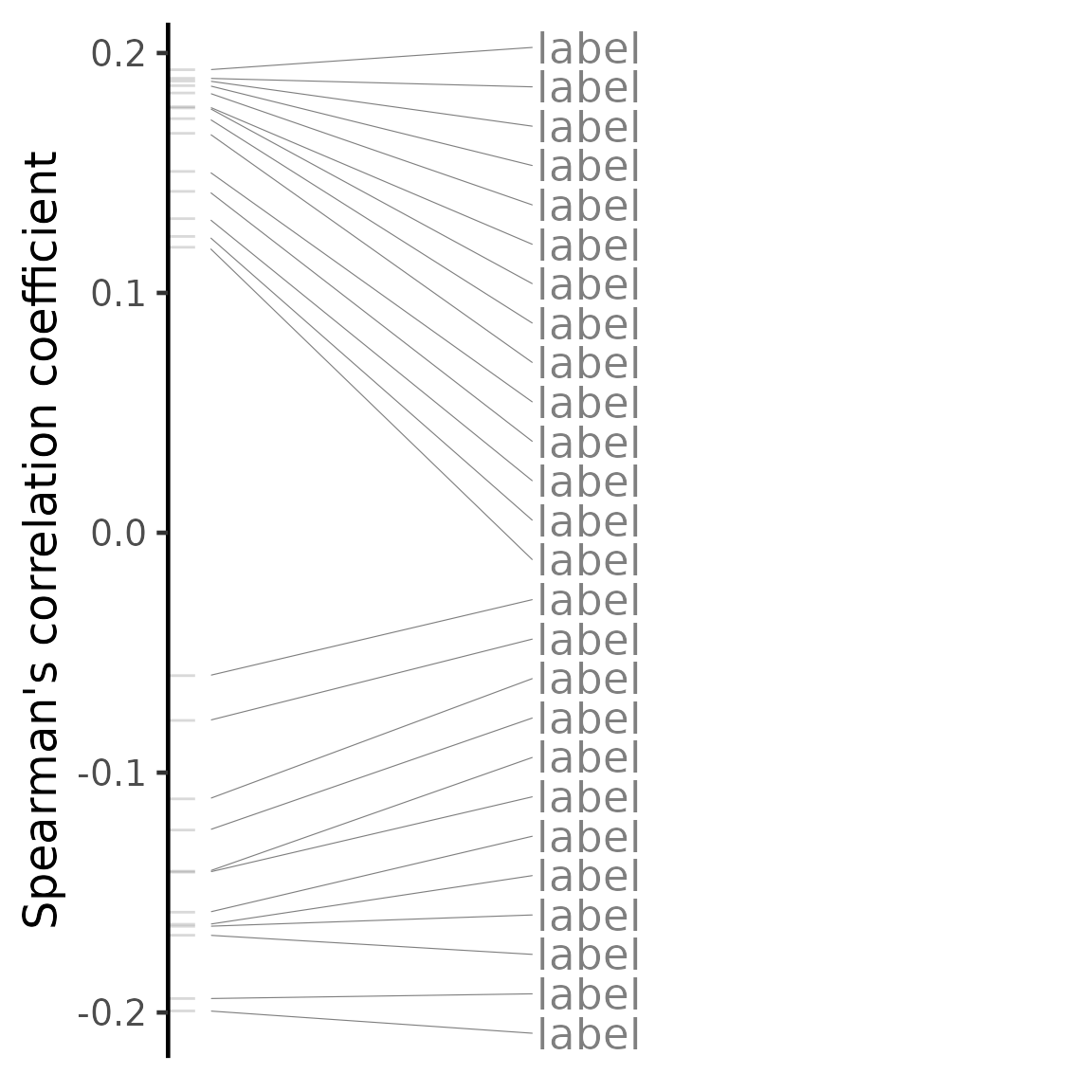
plot(compareKD, "pearson")## Warning: No shared levels found between `names(values)` of the manual scale and the
## data's colour values.
## No shared levels found between `names(values)` of the manual scale and the
## data's colour values.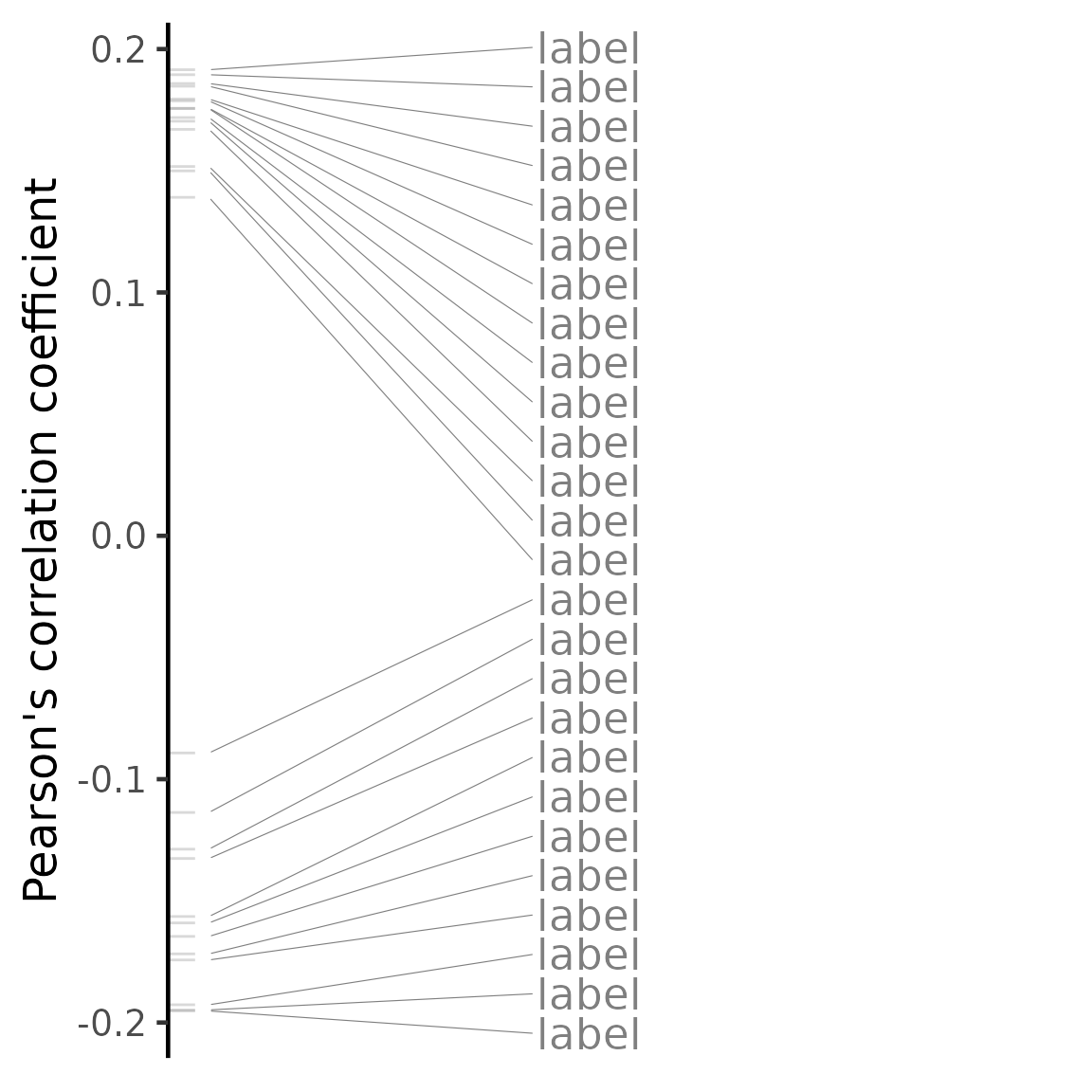
plot(compareKD, "gsea")## Warning: No shared levels found between `names(values)` of the manual scale and the
## data's colour values.
## No shared levels found between `names(values)` of the manual scale and the
## data's colour values.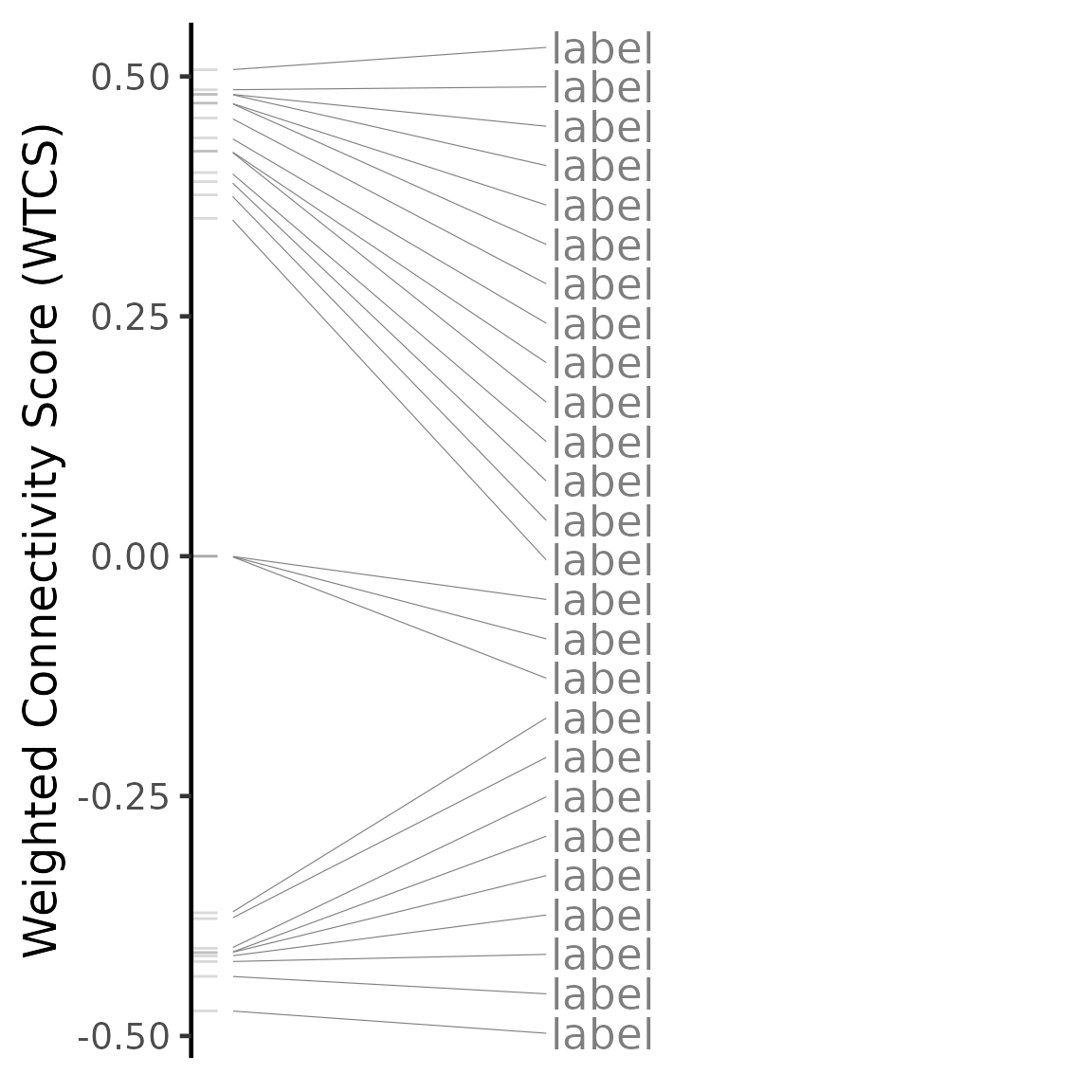
plot(compareKD, "rankProduct")## Warning: No shared levels found between `names(values)` of the manual scale and the
## data's colour values.
## No shared levels found between `names(values)` of the manual scale and the
## data's colour values.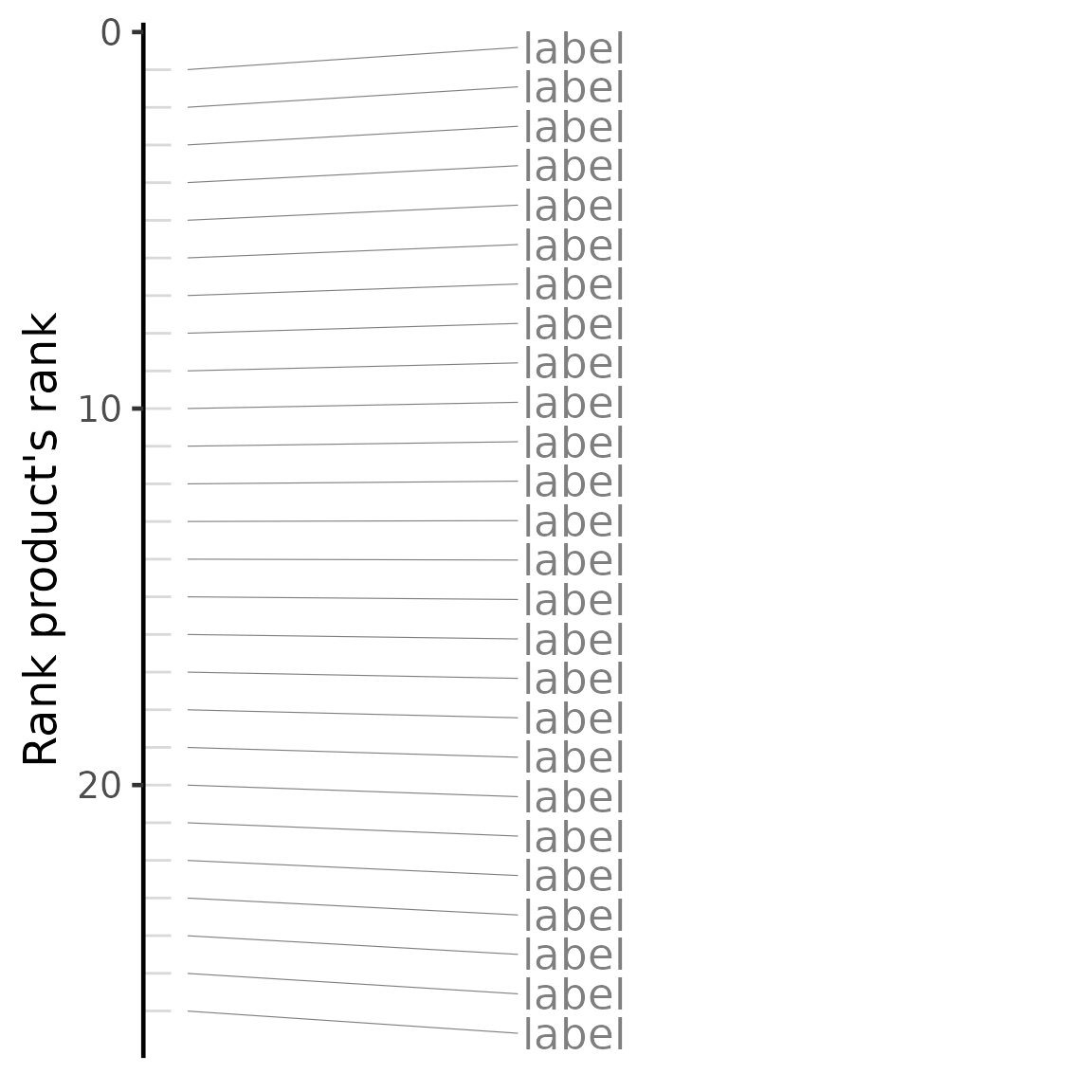
For small molecules:
# Most positively and negatively associated perturbations
compareCompounds[order(rankProduct_rank)]## compound_perturbation spearman_coef
## <char> <num>
## 1: CVD001_HEPG2_24H:BRD-A14014306-001-01-1:4.1 0.225335666
## 2: CVD001_HEPG2_24H:BRD-K84595254-001-03-0:4.9444 0.085133810
## 3: CVD001_HEPG2_24H:BRD-K62810658-001-05-6:4.6768 -0.050201852
## 4: CVD001_HEPG2_24H:BRD-K41172353-001-01-4:4.7 0.005161915
## 5: CVD001_HEPG2_24H:BRD-K60476892-001-02-1:4.1072 -0.077832390
## 6: CVD001_HEPG2_24H:BRD-K96188950-001-04-5:4.3967 -0.037174082
## 7: CVD001_HEPG2_24H:BRD-K84389640-001-01-5:4.225 -0.078331814
## 8: CVD001_HEPG2_24H:BRD-K77508012-001-01-9:6.025 -0.094208866
## 9: CVD001_HEPG2_24H:BRD-K31030218-001-01-1:4.25 -0.125531451
## 10: CVD001_HEPG2_24H:BRD-K94818765-001-01-0:4.8 -0.113418048
## 11: CVD001_HEPG2_24H:BRD-A65142661-034-01-8:5.35 -0.164686339
## spearman_pvalue spearman_qvalue pearson_coef pearson_pvalue pearson_qvalue
## <num> <num> <num> <num> <num>
## 1: 1.361508e-101 1.497659e-100 0.22640668 1.442316e-102 1.586548e-101
## 2: 1.294659e-15 2.373541e-15 0.06834399 1.415829e-10 2.224875e-10
## 3: 2.489663e-06 3.042921e-06 -0.02551843 1.673267e-02 1.840594e-02
## 4: 6.284641e-01 6.284641e-01 -0.01267514 2.347404e-01 2.347404e-01
## 5: 2.727173e-13 3.749863e-13 -0.06254059 4.400100e-09 6.050137e-09
## 6: 4.903648e-04 5.394013e-04 -0.04580625 1.737439e-05 2.123537e-05
## 7: 1.919930e-13 3.017033e-13 -0.08264025 8.485519e-15 1.555679e-14
## 8: 8.674931e-19 1.908485e-18 -0.08283071 7.364690e-15 1.555679e-14
## 9: 3.293334e-32 1.207556e-31 -0.12054061 8.215231e-30 3.012251e-29
## 10: 1.461350e-26 4.018714e-26 -0.11053910 2.640853e-25 7.262345e-25
## 11: 1.741438e-54 9.577908e-54 -0.14797060 3.242185e-44 1.783202e-43
## GSEA spearman_rank pearson_rank GSEA_rank rankProduct_rank
## <num> <num> <num> <num> <num>
## 1: 0.4932836 1 1 1.0 1
## 2: 0.2179143 2 2 3.0 2
## 3: 0.2304579 5 4 2.0 3
## 4: 0.0000000 3 3 5.5 4
## 5: 0.0000000 6 6 5.5 5
## 6: -0.2821337 4 5 10.0 6
## 7: 0.0000000 7 7 5.5 7
## 8: -0.2442628 8 8 8.0 8
## 9: 0.0000000 10 10 5.5 9
## 10: -0.2621164 9 9 9.0 10
## 11: -0.3381341 11 11 11.0 11
plot(compareCompounds, "rankProduct")## Warning: No shared levels found between `names(values)` of the manual scale and the
## data's colour values.
## No shared levels found between `names(values)` of the manual scale and the
## data's colour values.
The Gene Set Enrichment Analysis (GSEA) score is based on the Weighted Connectivity Score (WTCS), a composite and bi-directional version of the weighted Kolmogorov-Smirnov enrichment statistic (ES) (Subramanian et al., Cell 2017).
To calculate the GSEA score, GSEA is run for the most up- and down-regulated genes from the user’s differential expression profile. The GSEA score is the mean between EStop and ESbottom (however, if EStop and ESbottom have the same sign, the GSEA score is 0).
If a perturbation has a similar differential expression profile to our data (higher GSEA score), we expect to see the most up-regulated (down-regulated) genes in the perturbation enriched in the top (bottom) n differentially expressed genes from our data.
Information on perturbations
To get associated information as made available from CMap:
# Information on the EIF4G1 knockdown perturbation
EIF4G1knockdown <- grep("EIF4G1", compareKD[[1]], value=TRUE)
print(compareKD, EIF4G1knockdown)## $metadata
## Key: <sig_id, pert_id, pert_iname, pert_type, cell_id, pert_dose, pert_dose_unit, pert_idose, pert_time, pert_time_unit, pert_itime, distil_id>
## sig_id pert_id pert_iname pert_type cell_id
## <char> <char> <char> <char> <char>
## 1: CGS001_HEPG2_96H:EIF4G1:1.5 CGS001-1981 EIF4G1 trt_sh.cgs HEPG2
## pert_dose pert_dose_unit pert_idose pert_time pert_time_unit pert_itime
## <num> <char> <char> <int> <char> <char>
## 1: 1.5 microL 1.5 microL 96 h 96 h
## distil_id
## <char>
## 1: KDA004_HEPG2_96H:TRCN0000061769:-666|KDA004_HEPG2_96H:TRCN0000061770:-666|KDA004_HEPG2_96H:TRCN0000061772:-666
# Information on the top 10 most similar compound perturbations (based on
# Spearman's correlation)
print(compareKD[order(rankProduct_rank)], 1:10)## $metadata
## Key: <sig_id, pert_id, pert_iname, pert_type, cell_id, pert_dose, pert_dose_unit, pert_idose, pert_time, pert_time_unit, pert_itime, distil_id>
## sig_id pert_id pert_iname pert_type cell_id
## <char> <char> <char> <char> <char>
## 1: CGS001_HEPG2_96H:COPS5:1.5 CGS001-10987 COPS5 trt_sh.cgs HEPG2
## 2: CGS001_HEPG2_96H:EIF4G1:1.5 CGS001-1981 EIF4G1 trt_sh.cgs HEPG2
## 3: CGS001_HEPG2_96H:KIAA0196:1.5 CGS001-9897 KIAA0196 trt_sh.cgs HEPG2
## 4: CGS001_HEPG2_96H:KIF20A:1.5 CGS001-10112 KIF20A trt_sh.cgs HEPG2
## 5: CGS001_HEPG2_96H:MECP2:1.5 CGS001-4204 MECP2 trt_sh.cgs HEPG2
## 6: CGS001_HEPG2_96H:MEST:1.5 CGS001-4232 MEST trt_sh.cgs HEPG2
## 7: CGS001_HEPG2_96H:PPIH:1.5 CGS001-10465 PPIH trt_sh.cgs HEPG2
## 8: CGS001_HEPG2_96H:SKIV2L:1.5 CGS001-6499 SKIV2L trt_sh.cgs HEPG2
## 9: CGS001_HEPG2_96H:SQRDL:1.5 CGS001-58472 SQRDL trt_sh.cgs HEPG2
## 10: CGS001_HEPG2_96H:STAT1:1.5 CGS001-6772 STAT1 trt_sh.cgs HEPG2
## pert_dose pert_dose_unit pert_idose pert_time pert_time_unit pert_itime
## <num> <char> <char> <int> <char> <char>
## 1: 1.5 microL 1.5 microL 96 h 96 h
## 2: 1.5 microL 1.5 microL 96 h 96 h
## 3: 1.5 microL 1.5 microL 96 h 96 h
## 4: 1.5 microL 1.5 microL 96 h 96 h
## 5: 1.5 microL 1.5 microL 96 h 96 h
## 6: 1.5 microL 1.5 microL 96 h 96 h
## 7: 1.5 microL 1.5 microL 96 h 96 h
## 8: 1.5 microL 1.5 microL 96 h 96 h
## 9: 1.5 microL 1.5 microL 96 h 96 h
## 10: 1.5 microL 1.5 microL 96 h 96 h
## distil_id
## <char>
## 1: KDB009_HEPG2_96H:TRCN0000019200:-666|KDB009_HEPG2_96H:TRCN0000019201:-666|KDB009_HEPG2_96H:TRCN0000019202:-666
## 2: KDA004_HEPG2_96H:TRCN0000061769:-666|KDA004_HEPG2_96H:TRCN0000061770:-666|KDA004_HEPG2_96H:TRCN0000061772:-666
## 3: KDB010_HEPG2_96H:TRCN0000128018:-666|KDB010_HEPG2_96H:TRCN0000129686:-666|KDB010_HEPG2_96H:TRCN0000131227:-666
## 4: KDB009_HEPG2_96H:TRCN0000116523:-666|KDB009_HEPG2_96H:TRCN0000116524:-666|KDB009_HEPG2_96H:TRCN0000116525:-666
## 5: KDA003_HEPG2_96H:TRCN0000021241:-666|KDA003_HEPG2_96H:TRCN0000021242:-666
## 6: KDB008_HEPG2_96H:TRCN0000075318:-666|KDB008_HEPG2_96H:TRCN0000075320:-666|KDB008_HEPG2_96H:TRCN0000075321:-666
## 7: KDD006_HEPG2_96H:TRCN0000049389:-666|KDD006_HEPG2_96H:TRCN0000049390:-666
## 8: KDB008_HEPG2_96H:TRCN0000051813:-666|KDB008_HEPG2_96H:TRCN0000051815:-666|KDB008_HEPG2_96H:TRCN0000051816:-666
## 9: KDB009_HEPG2_96H:TRCN0000039004:-666|KDB009_HEPG2_96H:TRCN0000039005:-666|KDB009_HEPG2_96H:TRCN0000039007:-666
## 10: KDA005_HEPG2_96H:TRCN0000004265:-666|KDB007_HEPG2_96H:TRCN0000004265:-666|KDD006_HEPG2_96H:TRCN0000004265:-666|KDA005_HEPG2_96H:TRCN0000004266:-666|KDB007_HEPG2_96H:TRCN0000004266:-666|KDD006_HEPG2_96H:TRCN0000004266:-666|KDA005_HEPG2_96H:TRCN0000004267:-666|KDB007_HEPG2_96H:TRCN0000004267:-666
# Get table with all information available (including ranks, metadata, compound
# information, etc.)
table <- as.table(compareKD)
head(table)## gene_perturbation spearman_coef pearson_coef GSEA
## <char> <num> <num> <num>
## 1: CGS001_HEPG2_96H:EIF4G1:1.5 0.1770993 0.1915142 0.4861485
## 2: CGS001_HEPG2_96H:MECP2:1.5 0.1930335 0.1756788 0.3903862
## 3: CGS001_HEPG2_96H:SKIV2L:1.5 0.1423207 0.1755377 0.5070024
## 4: CGS001_HEPG2_96H:KIAA0196:1.5 0.1894301 0.1847352 0.3521034
## 5: CGS001_HEPG2_96H:STAT1:1.5 0.1833382 0.1894661 0.3765351
## 6: CGS001_HEPG2_96H:MEST:1.5 0.1726479 0.1858475 0.4219693
## spearman_rank pearson_rank GSEA_rank rankProduct_rank pert_id pert_iname
## <num> <num> <num> <num> <char> <char>
## 1: 7 1 2 1 CGS001-1981 EIF4G1
## 2: 1 7 12 2 CGS001-4204 MECP2
## 3: 11 8 1 3 CGS001-6499 SKIV2L
## 4: 2 4 14 4 CGS001-9897 KIAA0196
## 5: 5 2 13 5 CGS001-6772 STAT1
## 6: 8 3 10 6 CGS001-4232 MEST
## pert_type cell_id pert_idose pert_itime pr_gene_id
## <char> <char> <char> <char> <int>
## 1: trt_sh.cgs HEPG2 1.5 microL 96 h 1981
## 2: trt_sh.cgs HEPG2 1.5 microL 96 h 4204
## 3: trt_sh.cgs HEPG2 1.5 microL 96 h 6499
## 4: trt_sh.cgs HEPG2 1.5 microL 96 h 9897
## 5: trt_sh.cgs HEPG2 1.5 microL 96 h 6772
## 6: trt_sh.cgs HEPG2 1.5 microL 96 h 4232
## pr_gene_title pr_is_lm pr_is_bing
## <char> <int> <int>
## 1: eukaryotic translation initiation factor 4 gamma, 1 1 1
## 2: methyl-CpG binding protein 2 0 1
## 3: Ski2 like RNA helicase 1 1
## 4: KIAA0196 1 1
## 5: signal transducer and activator of transcription 1 1 1
## 6: mesoderm specific transcript 1 1
# Obtain available raw information from compared perturbations
names(attributes(compareKD)) # Data available in compared perturbations## [1] "class" "row.names" ".internal.selfref"
## [4] "names" "metadata" "geneInfo"
## [7] "input" "rankingInfo" "geneset"
## [10] "runtime"
attr(compareKD, "metadata") # Perturbation metadata## Key: <sig_id, pert_id, pert_iname, pert_type, cell_id, pert_dose, pert_dose_unit, pert_idose, pert_time, pert_time_unit, pert_itime, distil_id>
## sig_id pert_id pert_iname pert_type cell_id
## <char> <char> <char> <char> <char>
## 1: CGS001_HEPG2_96H:CDCA8:1.5 CGS001-55143 CDCA8 trt_sh.cgs HEPG2
## 2: CGS001_HEPG2_96H:COPS5:1.5 CGS001-10987 COPS5 trt_sh.cgs HEPG2
## 3: CGS001_HEPG2_96H:DHX16:1.5 CGS001-8449 DHX16 trt_sh.cgs HEPG2
## 4: CGS001_HEPG2_96H:EHMT2:1.5 CGS001-10919 EHMT2 trt_sh.cgs HEPG2
## 5: CGS001_HEPG2_96H:EIF4G1:1.5 CGS001-1981 EIF4G1 trt_sh.cgs HEPG2
## 6: CGS001_HEPG2_96H:EYA1:1.5 CGS001-2138 EYA1 trt_sh.cgs HEPG2
## 7: CGS001_HEPG2_96H:GNAI2:1.5 CGS001-2771 GNAI2 trt_sh.cgs HEPG2
## 8: CGS001_HEPG2_96H:GTPBP8:1.5 CGS001-29083 GTPBP8 trt_sh.cgs HEPG2
## 9: CGS001_HEPG2_96H:HFE:1.5 CGS001-3077 HFE trt_sh.cgs HEPG2
## 10: CGS001_HEPG2_96H:KIAA0196:1.5 CGS001-9897 KIAA0196 trt_sh.cgs HEPG2
## 11: CGS001_HEPG2_96H:KIF20A:1.5 CGS001-10112 KIF20A trt_sh.cgs HEPG2
## 12: CGS001_HEPG2_96H:MAF:1.5 CGS001-4094 MAF trt_sh.cgs HEPG2
## 13: CGS001_HEPG2_96H:MECP2:1.5 CGS001-4204 MECP2 trt_sh.cgs HEPG2
## 14: CGS001_HEPG2_96H:MEST:1.5 CGS001-4232 MEST trt_sh.cgs HEPG2
## 15: CGS001_HEPG2_96H:NDUFB6:1.5 CGS001-4712 NDUFB6 trt_sh.cgs HEPG2
## 16: CGS001_HEPG2_96H:PLOD2:1.5 CGS001-5352 PLOD2 trt_sh.cgs HEPG2
## 17: CGS001_HEPG2_96H:PPIH:1.5 CGS001-10465 PPIH trt_sh.cgs HEPG2
## 18: CGS001_HEPG2_96H:SHB:1.5 CGS001-6461 SHB trt_sh.cgs HEPG2
## 19: CGS001_HEPG2_96H:SIAH2:1.5 CGS001-6478 SIAH2 trt_sh.cgs HEPG2
## 20: CGS001_HEPG2_96H:SKIV2L:1.5 CGS001-6499 SKIV2L trt_sh.cgs HEPG2
## 21: CGS001_HEPG2_96H:SQRDL:1.5 CGS001-58472 SQRDL trt_sh.cgs HEPG2
## 22: CGS001_HEPG2_96H:STAT1:1.5 CGS001-6772 STAT1 trt_sh.cgs HEPG2
## 23: CGS001_HEPG2_96H:SULT1A2:1.5 CGS001-6799 SULT1A2 trt_sh.cgs HEPG2
## 24: CGS001_HEPG2_96H:TMEM5:1.5 CGS001-10329 TMEM5 trt_sh.cgs HEPG2
## 25: CGS001_HEPG2_96H:UBAP2L:1.5 CGS001-9898 UBAP2L trt_sh.cgs HEPG2
## 26: CGS001_HEPG2_96H:ZBTB24:1.5 CGS001-9841 ZBTB24 trt_sh.cgs HEPG2
## sig_id pert_id pert_iname pert_type cell_id
## <char> <char> <char> <char> <char>
## pert_dose pert_dose_unit pert_idose pert_time pert_time_unit pert_itime
## <num> <char> <char> <int> <char> <char>
## 1: 1.5 microL 1.5 microL 96 h 96 h
## 2: 1.5 microL 1.5 microL 96 h 96 h
## 3: 1.5 microL 1.5 microL 96 h 96 h
## 4: 1.5 microL 1.5 microL 96 h 96 h
## 5: 1.5 microL 1.5 microL 96 h 96 h
## 6: 1.5 microL 1.5 microL 96 h 96 h
## 7: 1.5 microL 1.5 microL 96 h 96 h
## 8: 1.5 microL 1.5 microL 96 h 96 h
## 9: 1.5 microL 1.5 microL 96 h 96 h
## 10: 1.5 microL 1.5 microL 96 h 96 h
## 11: 1.5 microL 1.5 microL 96 h 96 h
## 12: 1.5 microL 1.5 microL 96 h 96 h
## 13: 1.5 microL 1.5 microL 96 h 96 h
## 14: 1.5 microL 1.5 microL 96 h 96 h
## 15: 1.5 microL 1.5 microL 96 h 96 h
## 16: 1.5 microL 1.5 microL 96 h 96 h
## 17: 1.5 microL 1.5 microL 96 h 96 h
## 18: 1.5 microL 1.5 microL 96 h 96 h
## 19: 1.5 microL 1.5 microL 96 h 96 h
## 20: 1.5 microL 1.5 microL 96 h 96 h
## 21: 1.5 microL 1.5 microL 96 h 96 h
## 22: 1.5 microL 1.5 microL 96 h 96 h
## 23: 1.5 microL 1.5 microL 96 h 96 h
## 24: 1.5 microL 1.5 microL 96 h 96 h
## 25: 1.5 microL 1.5 microL 96 h 96 h
## 26: 1.5 microL 1.5 microL 96 h 96 h
## pert_dose pert_dose_unit pert_idose pert_time pert_time_unit pert_itime
## <num> <char> <char> <int> <char> <char>
## distil_id
## <char>
## 1: KDA003_HEPG2_96H:TRCN0000007898:-666|KDA003_HEPG2_96H:TRCN0000007899:-666|KDA003_HEPG2_96H:TRCN0000007900:-666
## 2: KDB009_HEPG2_96H:TRCN0000019200:-666|KDB009_HEPG2_96H:TRCN0000019201:-666|KDB009_HEPG2_96H:TRCN0000019202:-666
## 3: KDB010_HEPG2_96H:TRCN0000001123:-666|KDB010_HEPG2_96H:TRCN0000001124:-666|KDB010_HEPG2_96H:TRCN0000001125:-666
## 4: KDA002_HEPG2_96H:TRCN0000115667:-666|KDA002_HEPG2_96H:TRCN0000115669:-666|KDA002_HEPG2_96H:TRCN0000115671:-666
## 5: KDA004_HEPG2_96H:TRCN0000061769:-666|KDA004_HEPG2_96H:TRCN0000061770:-666|KDA004_HEPG2_96H:TRCN0000061772:-666
## 6: KDC008_HEPG2_96H:TRCN0000083443:-666|KDC008_HEPG2_96H:TRCN0000083446:-666|KDC008_HEPG2_96H:TRCN0000303461:-666
## 7: KDA006_HEPG2_96H:TRCN0000029514:-666|KDB003_HEPG2_96H:TRCN0000029514:-666|KDA006_HEPG2_96H:TRCN0000029517:-666|KDB003_HEPG2_96H:TRCN0000029517:-666|KDA006_HEPG2_96H:TRCN0000278333:-666|KDB003_HEPG2_96H:TRCN0000029518:-666
## 8: KDB001_HEPG2_96H:TRCN0000343725:-666|KDB001_HEPG2_96H:TRCN0000343727:-666|KDB001_HEPG2_96H:TRCN0000343728:-666
## 9: KDB008_HEPG2_96H:TRCN0000060018:-666|KDB008_HEPG2_96H:TRCN0000060019:-666|KDB008_HEPG2_96H:TRCN0000060022:-666
## 10: KDB010_HEPG2_96H:TRCN0000128018:-666|KDB010_HEPG2_96H:TRCN0000129686:-666|KDB010_HEPG2_96H:TRCN0000131227:-666
## 11: KDB009_HEPG2_96H:TRCN0000116523:-666|KDB009_HEPG2_96H:TRCN0000116524:-666|KDB009_HEPG2_96H:TRCN0000116525:-666
## 12: KDC001_HEPG2_96H:TRCN0000000254:-666|KDC001_HEPG2_96H:TRCN0000000256:-666|KDC001_HEPG2_96H:TRCN0000000258:-666
## 13: KDA003_HEPG2_96H:TRCN0000021241:-666|KDA003_HEPG2_96H:TRCN0000021242:-666
## 14: KDB008_HEPG2_96H:TRCN0000075318:-666|KDB008_HEPG2_96H:TRCN0000075320:-666|KDB008_HEPG2_96H:TRCN0000075321:-666
## 15: KDB010_HEPG2_96H:TRCN0000028409:-666|KDB010_HEPG2_96H:TRCN0000028417:-666|KDB010_HEPG2_96H:TRCN0000028428:-666
## 16: KDC010_HEPG2_96H:TRCN0000064808:-666|KDC010_HEPG2_96H:TRCN0000064809:-666|KDC010_HEPG2_96H:TRCN0000064810:-666
## 17: KDD006_HEPG2_96H:TRCN0000049389:-666|KDD006_HEPG2_96H:TRCN0000049390:-666
## 18: KDB001_HEPG2_96H:TRCN0000149080:-666|KDB001_HEPG2_96H:TRCN0000282026:-666|KDB001_HEPG2_96H:TRCN0000285376:-666
## 19: KDA005_HEPG2_96H:TRCN0000007415:-666|KDA005_HEPG2_96H:TRCN0000007416:-666|KDA005_HEPG2_96H:TRCN0000007418:-666
## 20: KDB008_HEPG2_96H:TRCN0000051813:-666|KDB008_HEPG2_96H:TRCN0000051815:-666|KDB008_HEPG2_96H:TRCN0000051816:-666
## 21: KDB009_HEPG2_96H:TRCN0000039004:-666|KDB009_HEPG2_96H:TRCN0000039005:-666|KDB009_HEPG2_96H:TRCN0000039007:-666
## 22: KDA005_HEPG2_96H:TRCN0000004265:-666|KDB007_HEPG2_96H:TRCN0000004265:-666|KDD006_HEPG2_96H:TRCN0000004265:-666|KDA005_HEPG2_96H:TRCN0000004266:-666|KDB007_HEPG2_96H:TRCN0000004266:-666|KDD006_HEPG2_96H:TRCN0000004266:-666|KDA005_HEPG2_96H:TRCN0000004267:-666|KDB007_HEPG2_96H:TRCN0000004267:-666
## 23: KDA001_HEPG2_96H:TRCN0000035200:-666|KDA001_HEPG2_96H:TRCN0000035202:-666|KDA001_HEPG2_96H:TRCN0000035203:-666
## 24: KDB001_HEPG2_96H:TRCN0000135504:-666|KDB001_HEPG2_96H:TRCN0000278433:-666
## 25: KDA003_HEPG2_96H:TRCN0000007677:-666|KDA003_HEPG2_96H:TRCN0000007680:-666|KDA003_HEPG2_96H:TRCN0000007681:-666
## 26: KDB008_HEPG2_96H:TRCN0000107435:-666|KDB008_HEPG2_96H:TRCN0000107437:-666|KDB008_HEPG2_96H:TRCN0000107438:-666
## distil_id
## <char>
attr(compareKD, "geneInfo") # Gene information## pr_gene_id pr_gene_symbol
## <int> <char>
## 1: 780 DDR1
## 2: 7849 PAX8
## 3: 2978 GUCA1A
## 4: 2049 EPHB3
## 5: 2101 ESRRA
## ---
## 12324: 4034 LRCH4
## 12325: 399664 MEX3D
## 12326: 54869 EPS8L1
## 12327: 90379 DCAF15
## 12328: 60 ACTB
## pr_gene_title
## <char>
## 1: discoidin domain receptor tyrosine kinase 1
## 2: paired box 8
## 3: guanylate cyclase activator 1A
## 4: EPH receptor B3
## 5: estrogen related receptor alpha
## ---
## 12324: leucine-rich repeats and calponin homology (CH) domain containing 4
## 12325: mex-3 RNA binding family member D
## 12326: EPS8 like 1
## 12327: DDB1 and CUL4 associated factor 15
## 12328: actin, beta
## pr_is_lm pr_is_bing
## <int> <int>
## 1: 1 1
## 2: 1 1
## 3: 0 0
## 4: 0 1
## 5: 0 1
## ---
## 12324: 0 1
## 12325: 0 1
## 12326: 0 1
## 12327: 0 1
## 12328: 0 1Relationship plots
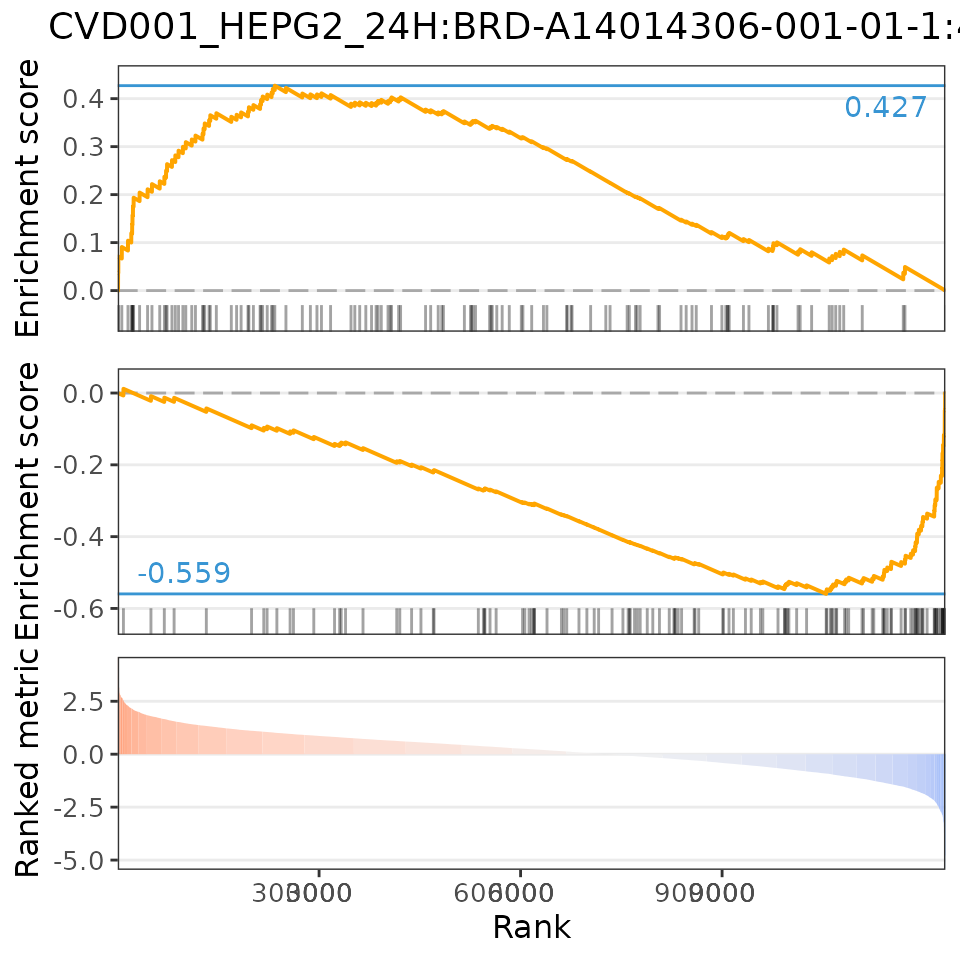
To analyse the relationship between the user-provided differential expression profile with that associated with a specific perturbation, scatter plots (for Spearman and Pearson analyses) and GSEA plots are available.
For instance, let’s plot the relationship between EIF4G1 shRNA knockdown from ENCODE with the CMap knockdown perturbations:
# Plot relationship with EIF4G1 knockdown from CMap
plot(compareKD, EIF4G1knockdown, "spearman")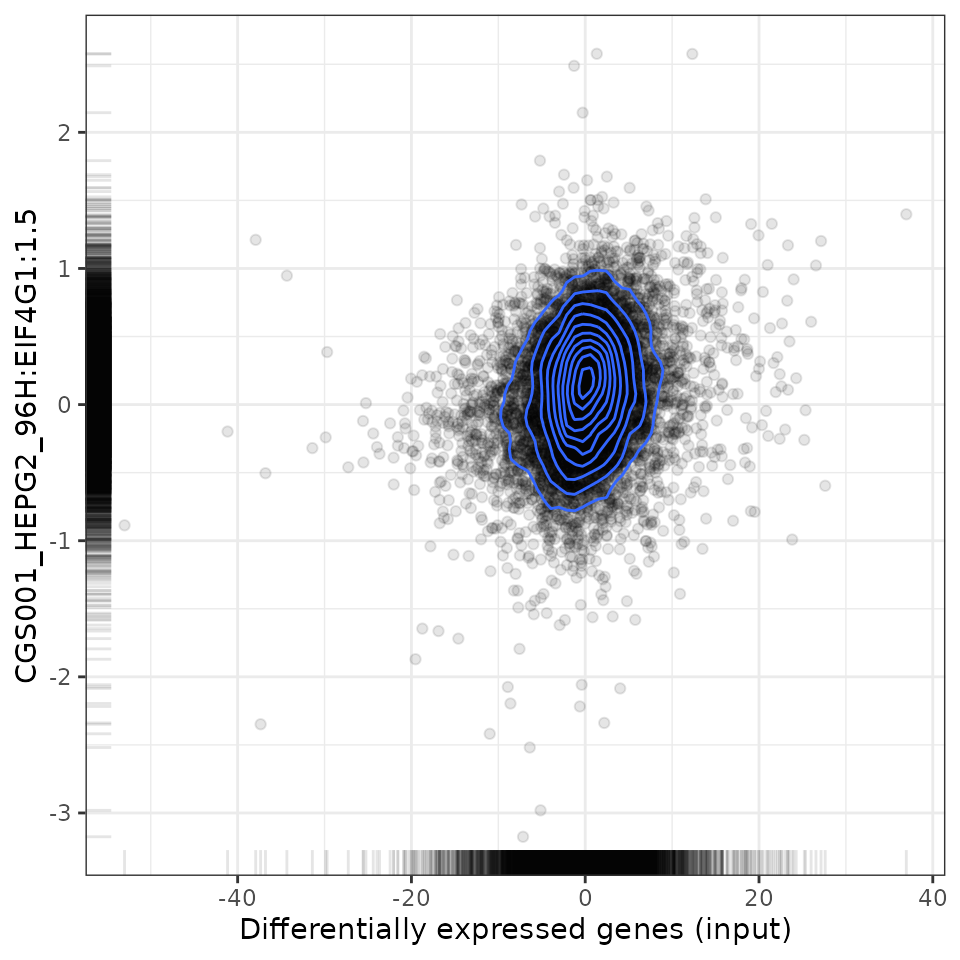
plot(compareKD, EIF4G1knockdown, "pearson")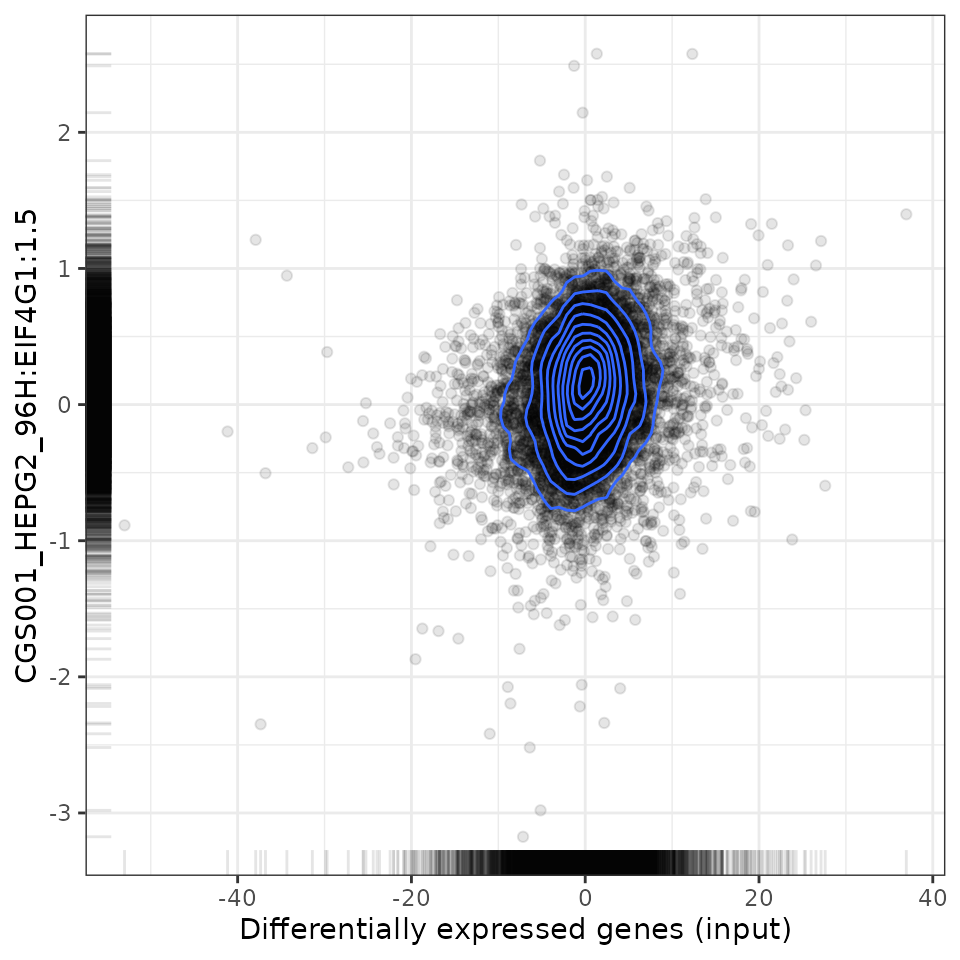
plot(compareKD, EIF4G1knockdown, "gsea")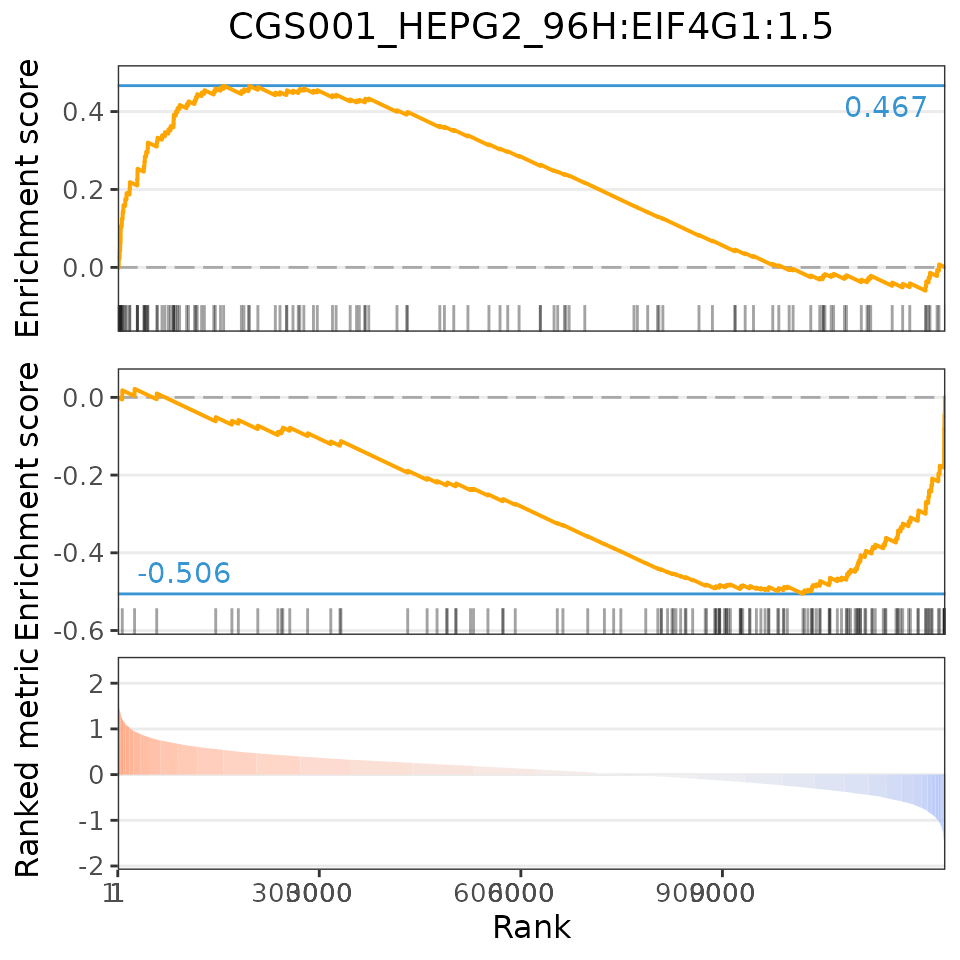
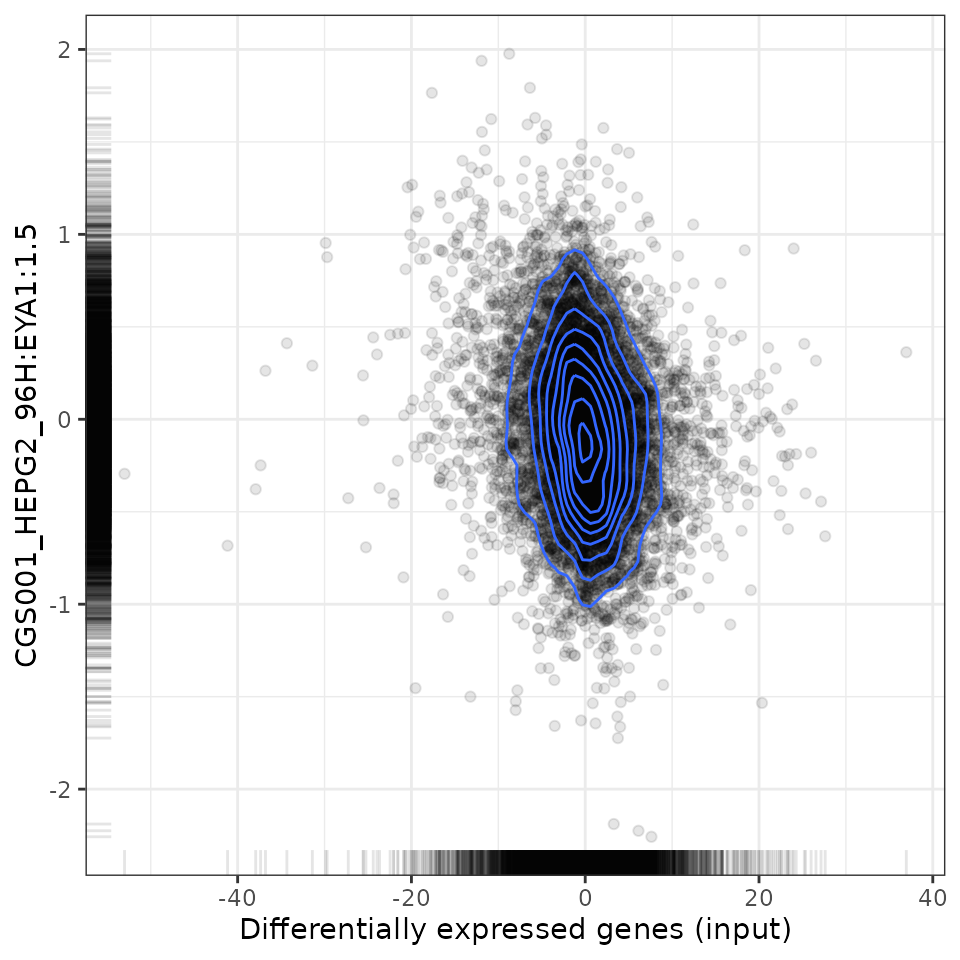
# Plot relationship with most negatively associated perturbation
plot(compareKD, compareKD[order(-spearman_rank)][1, 1], "spearman")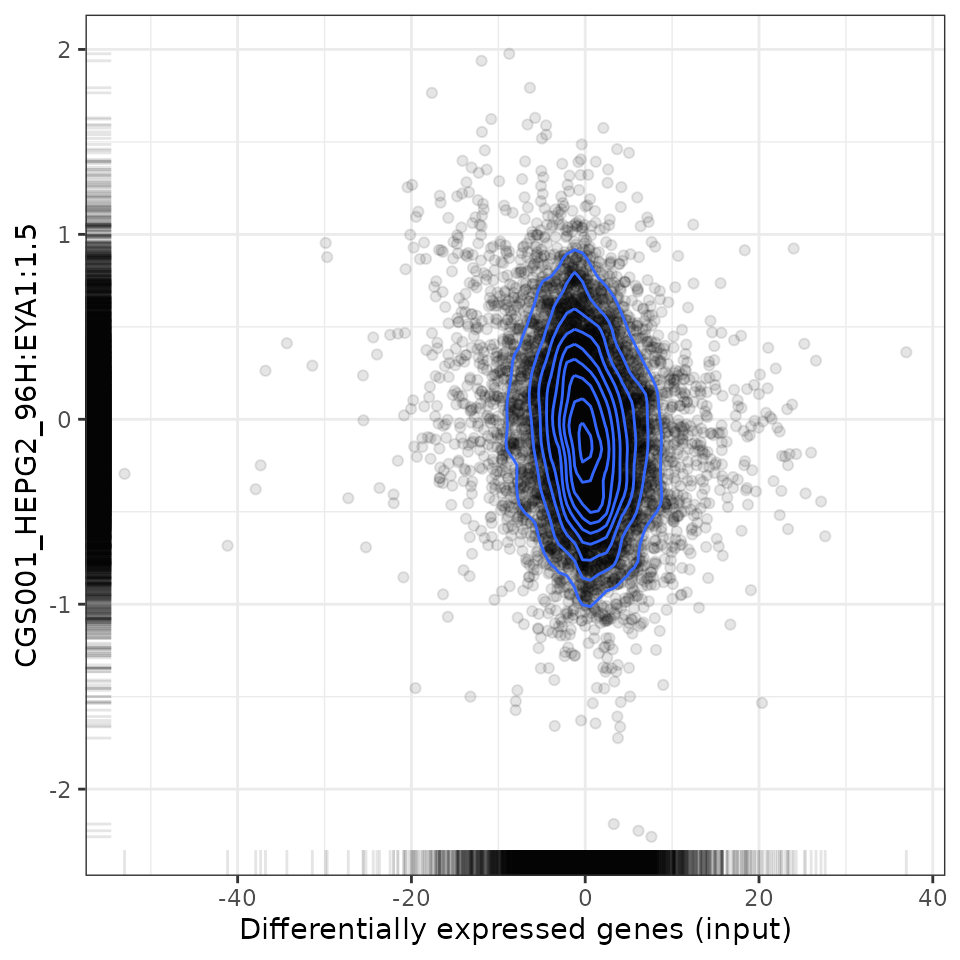
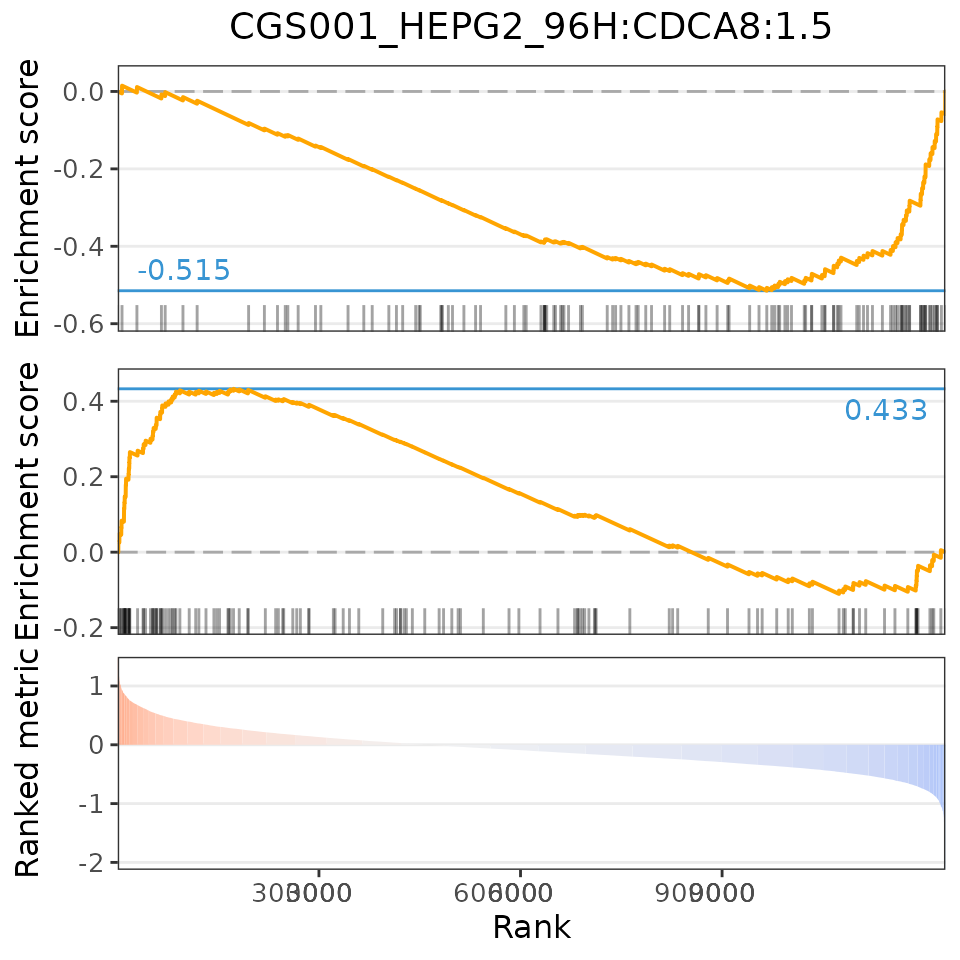
For small molecules:
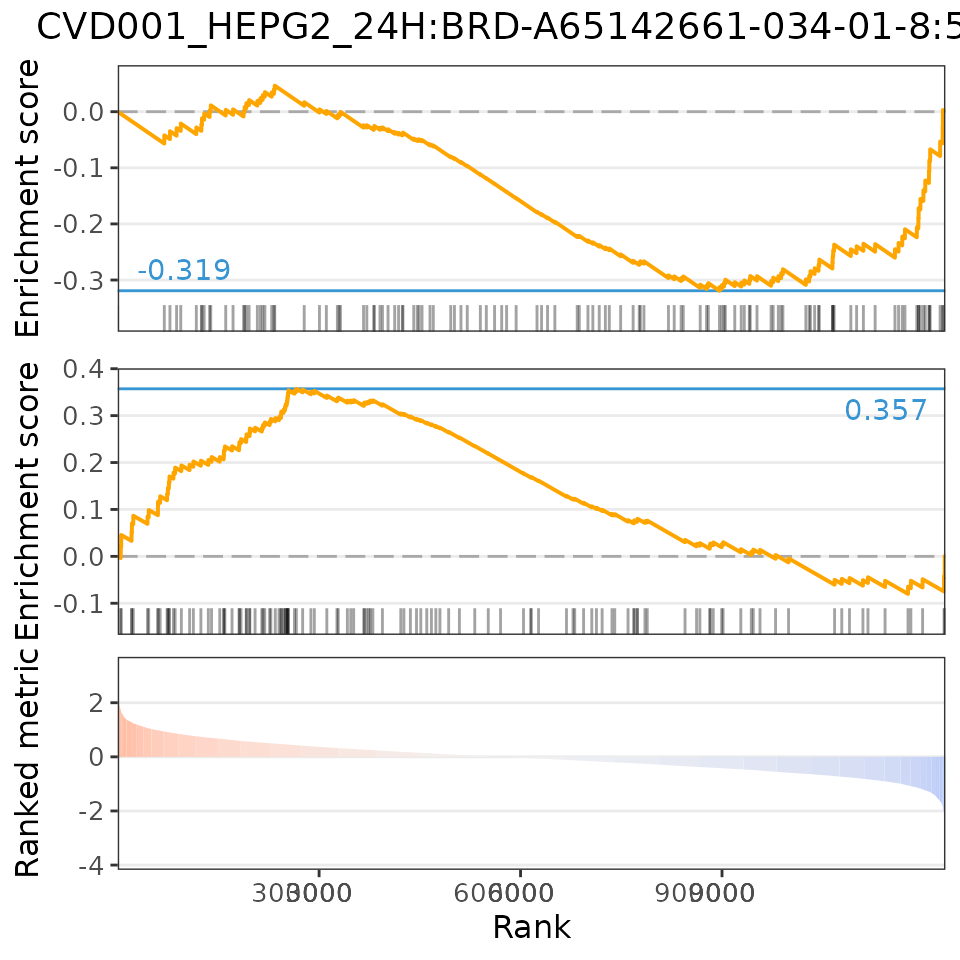
# Plot relationship with most positively associated perturbation
plot(compareCompounds, compareCompounds[order(spearman_rank)][1, 1], "spearman")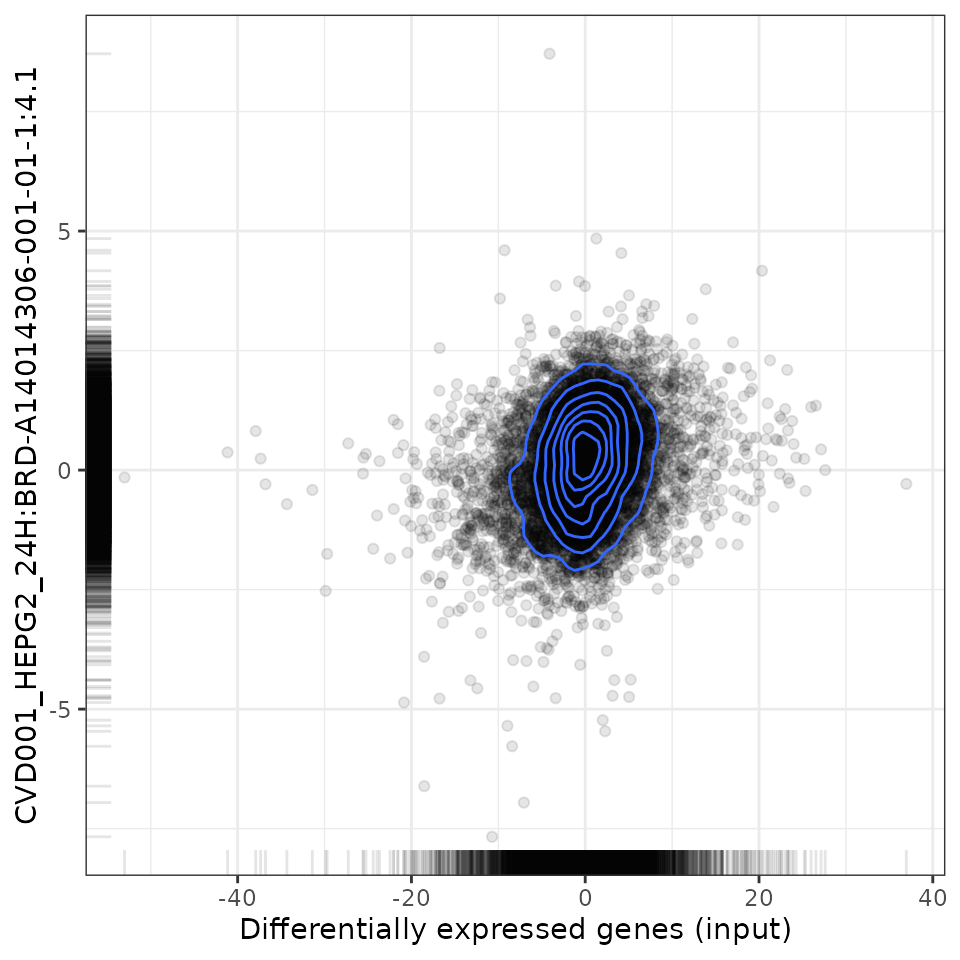
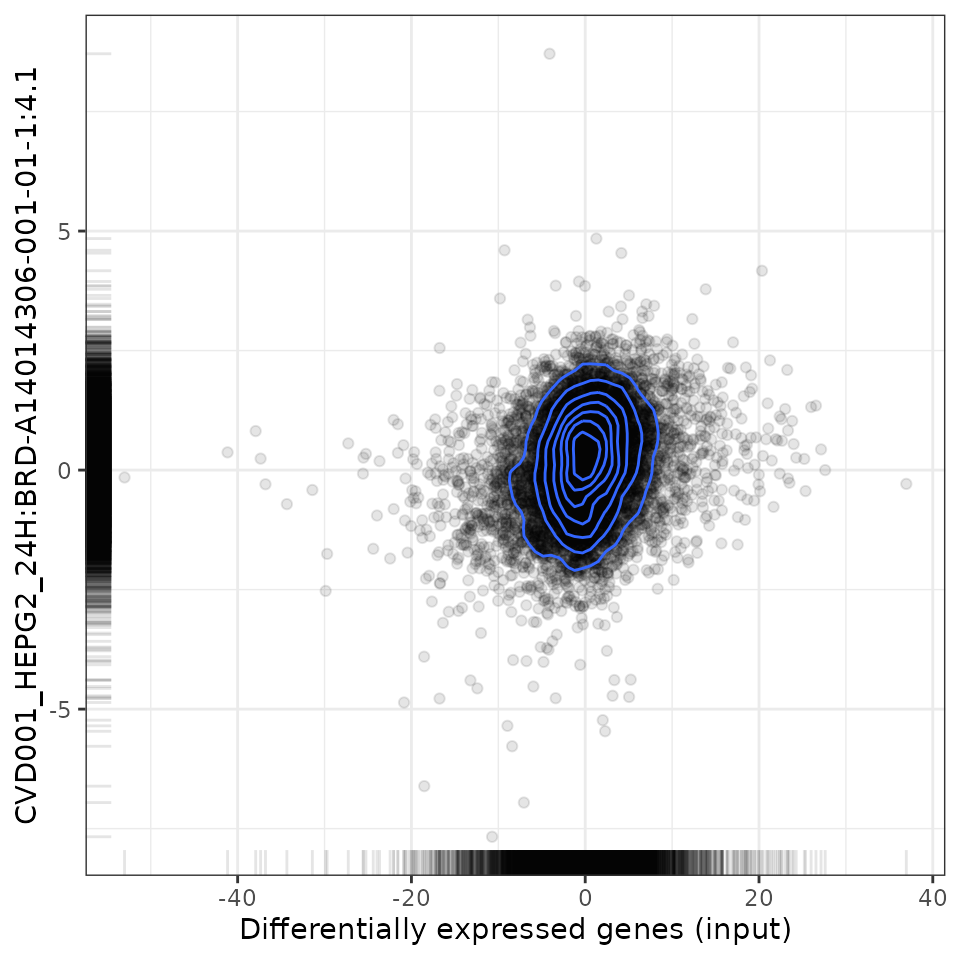
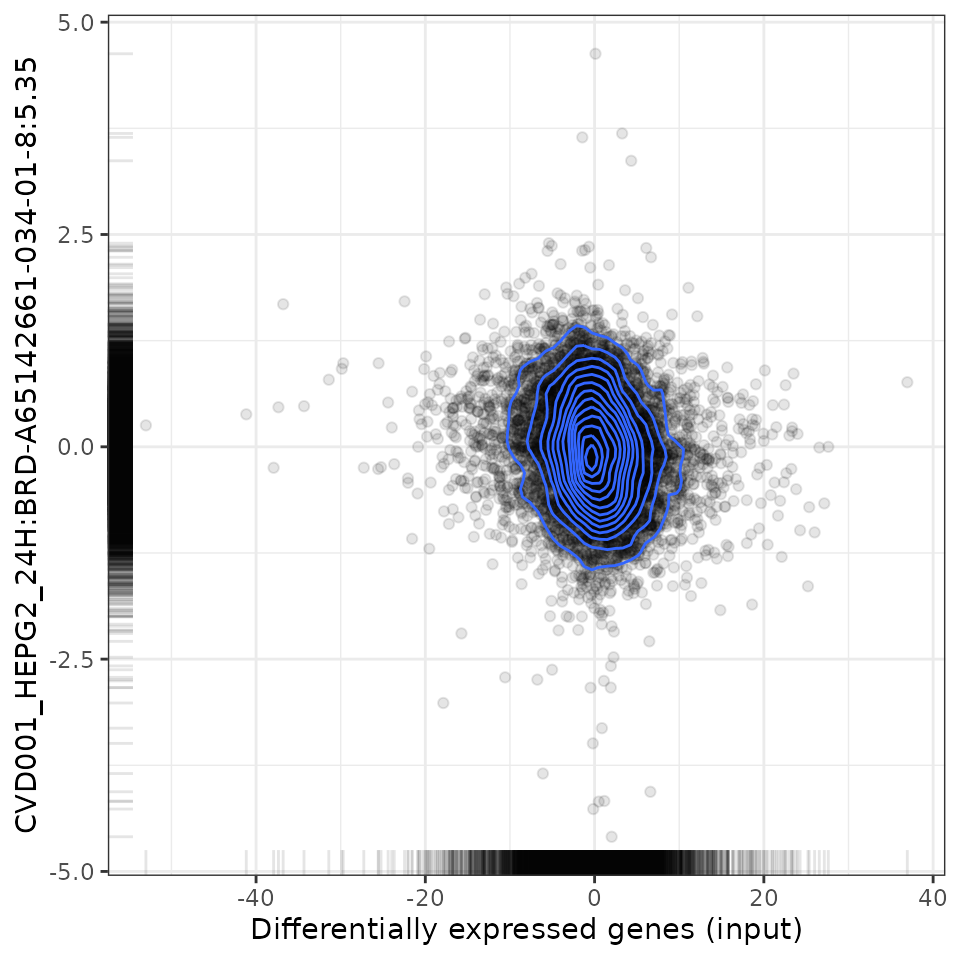
# Plot relationship with most negatively associated perturbation
plot(compareCompounds, compareCompounds[order(-spearman_rank)][1,1], "spearman")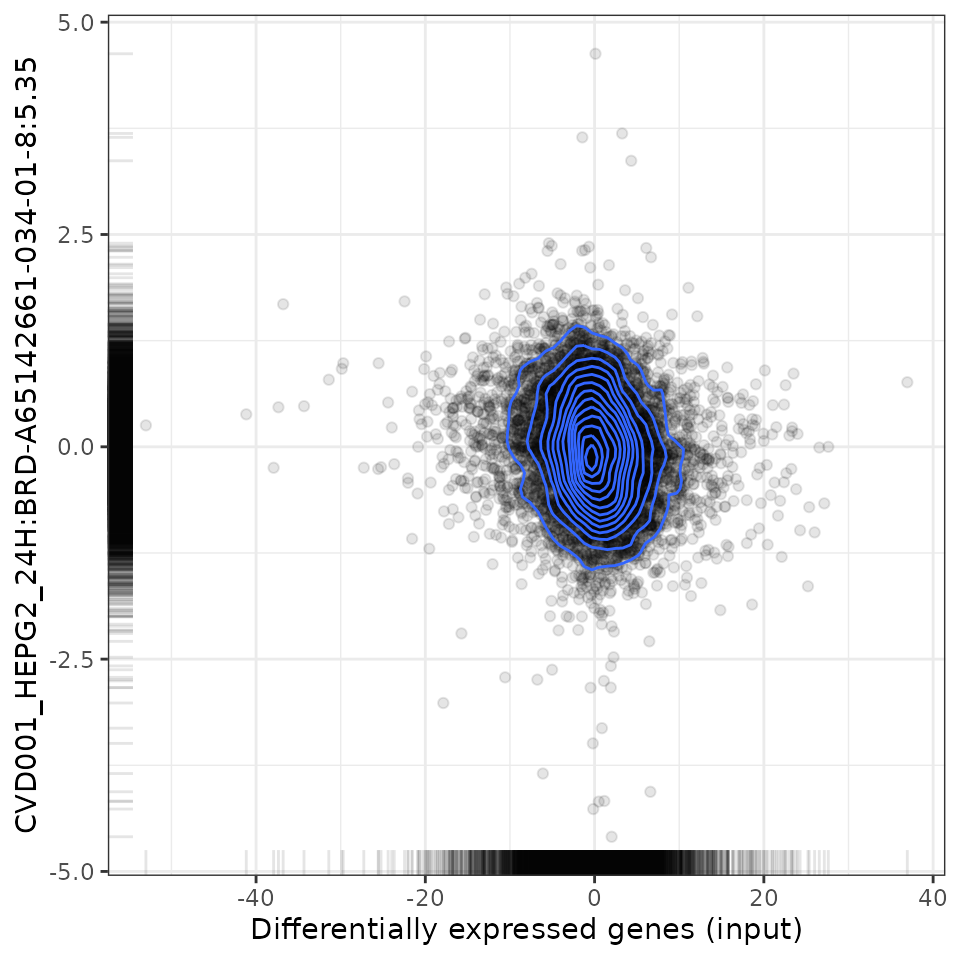
Predict targeting drugs
Compounds that target the phenotypes associated with the user-provided differential expression profile can be inferred by comparing against gene expression and drug sensitivity associations. The gene expression and drug sensitivity datasets derived from the following sources were correlated using Spearman’s correlation across the available cell lines.
| Source | Screened compounds | Human cancer cell lines |
|---|---|---|
| NCI60 | > 100 000 | 60 |
| GDSC 7 | 481 | 860 |
| CTRP 2.1 | 138 | ~700 |
To use an expression and drug sensitivity association based on
CTRP 2.1 (GDSC 7 and NCI60 could
be used instead) to infer targeting drugs for the user’s differential
expression profile:
## [1] "GDSC 7" "CTRP 2.1" "NCI60"
ctrp <- listExpressionDrugSensitivityAssociation()[[2]]
assoc <- loadExpressionDrugSensitivityAssociation(ctrp)## expressionDrugSensitivityCorCTRP2.1.qs not found: downloading data...## Loading data from expressionDrugSensitivityCorCTRP2.1.qs...
predicted <- predictTargetingDrugs(diffExprStat, assoc)## Subsetting data based on 11227 intersecting genes (83% of the 13451 input genes)...## Comparing against 545 CTRP 2.1 compounds (823 cell lines) using 'spearman, pearson, gsea' (gene size of 150)...## Comparison performed in 6.21 secs
plot(predicted, method="rankProduct")## Warning: No shared levels found between `names(values)` of the manual scale and the
## data's colour values.
## No shared levels found between `names(values)` of the manual scale and the
## data's colour values.## Warning: ggrepel: 535 unlabeled data points (too many overlaps). Consider
## increasing max.overlaps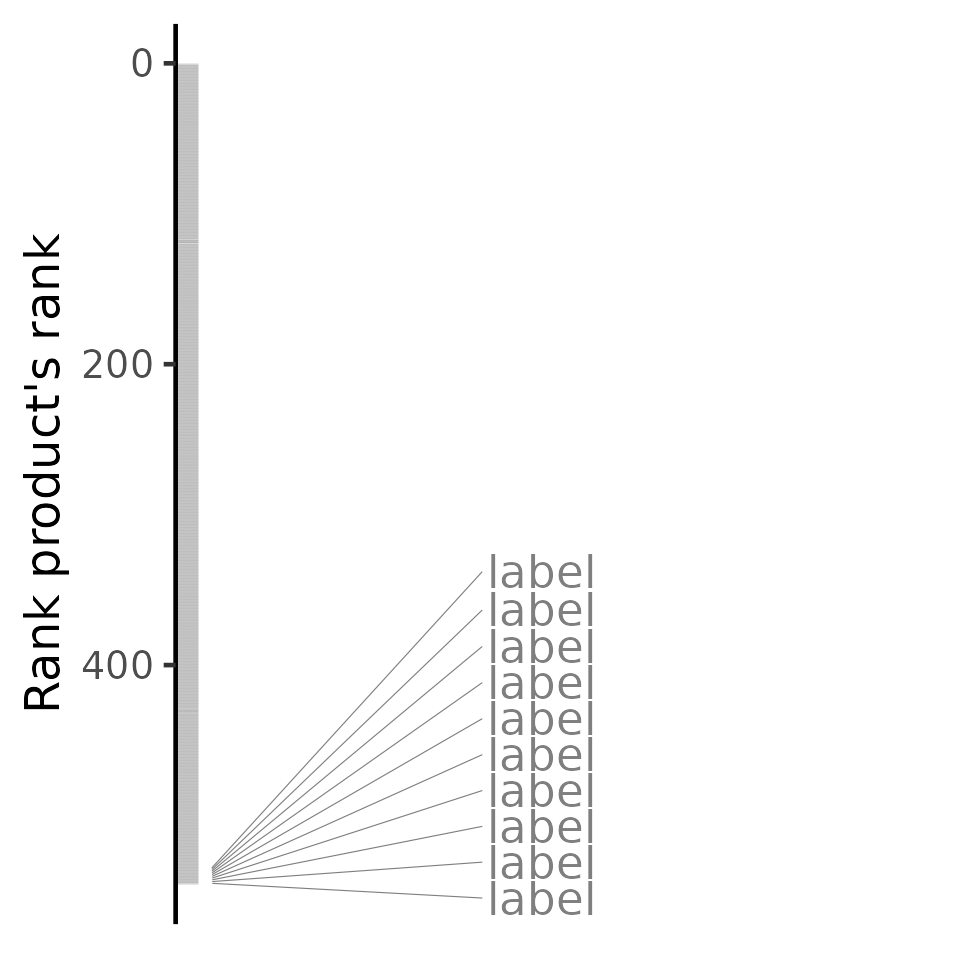
# Plot results for a given drug
plot(predicted, predicted[[1, 1]], method="spearman")## Loading data from /home/runner/work/cTRAP/cTRAP/vignettes/expressionDrugSensitivityCorCTRP2.1.qs...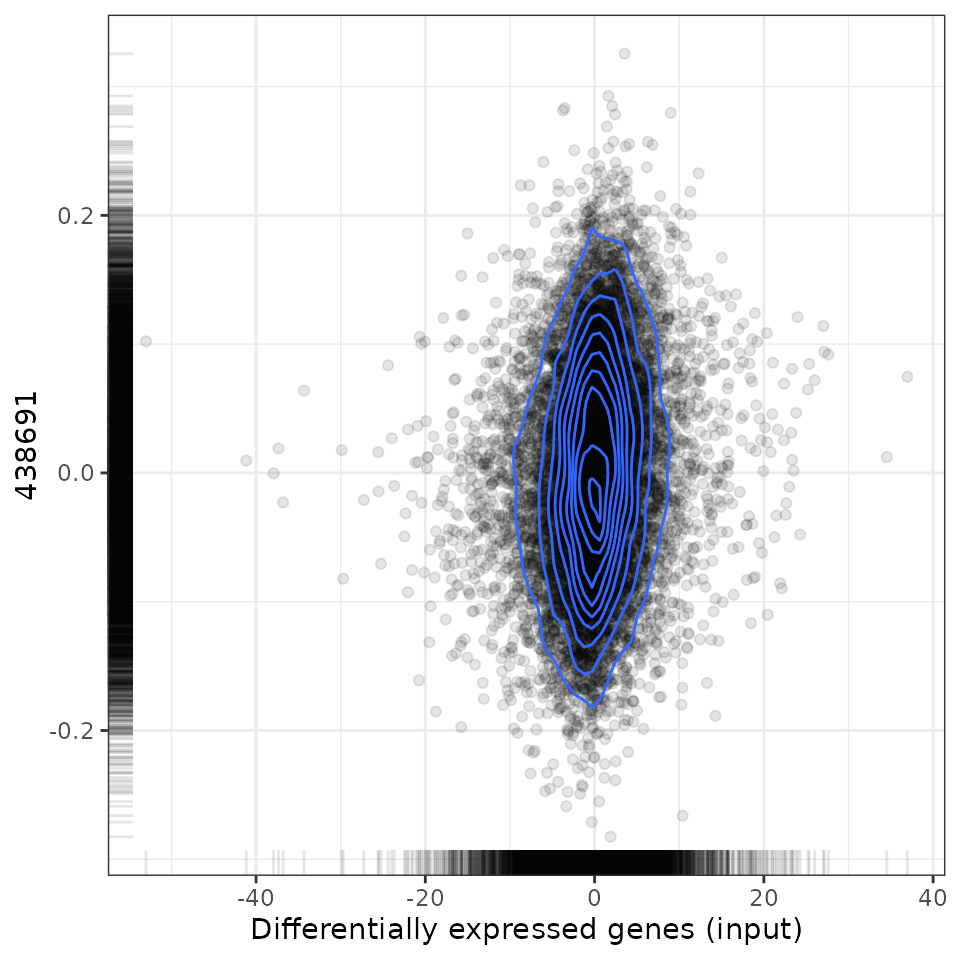
plot(predicted, predicted[[1, 1]], method="gsea")## Loading data from /home/runner/work/cTRAP/cTRAP/vignettes/expressionDrugSensitivityCorCTRP2.1.qs...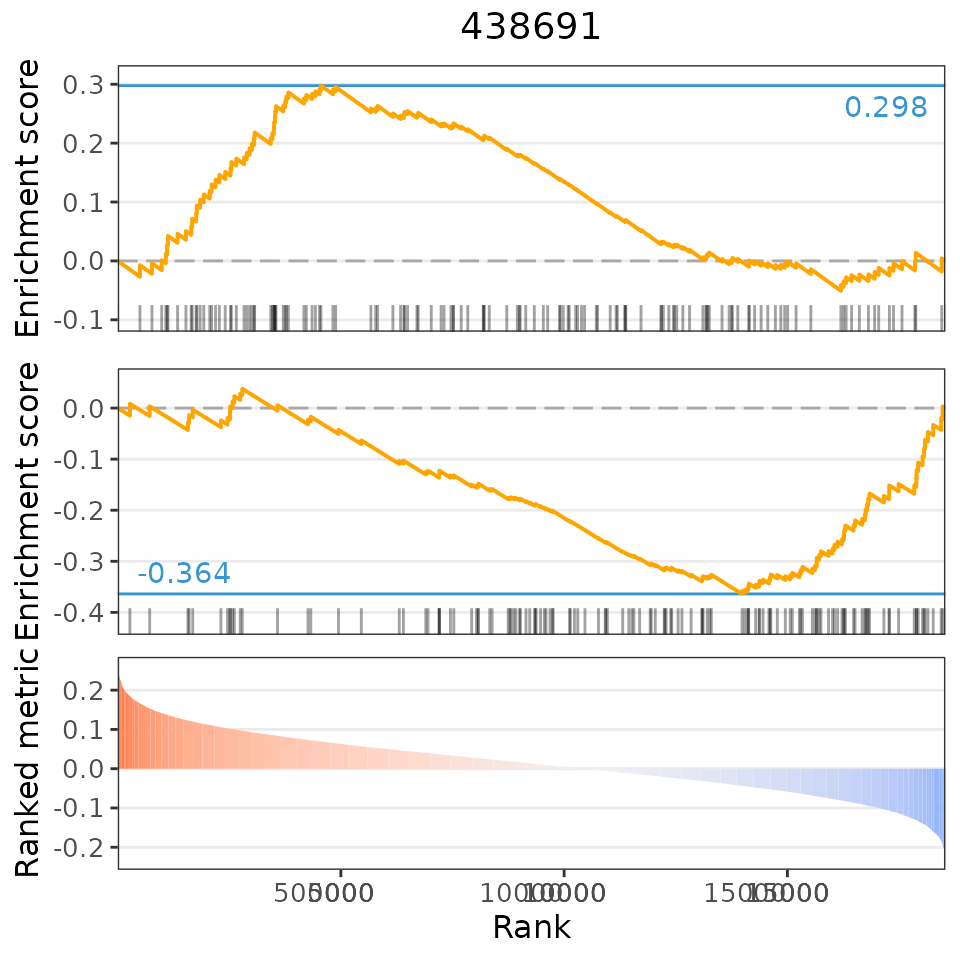
Compounds are ranked by their relative targeting potential based on the input differential expression profile (i.e. the 1st-ranked compound has higher targeting potential than the 2nd-ranked one).
Candidate targeting drugs can be plotted against the similarity ranking of their perturbations towards the user’s differential expression profile. Note that the highlighted values are the same compounds for the following plots annotated with their name, gene target and mechanism of action (MOA), respectively.
# Label by compound name
plotTargetingDrugsVSsimilarPerturbations(
predicted, compareCompounds, column="spearman_rank")## Columns 'broad id' and 'pubchem_cid' were matched based on 0 common values; to manually select columns to compare, please set arguments starting with 'keyCol'## Warning: No shared levels found between `names(values)` of the manual scale and the
## data's colour values.
# Label by compound's gene target
plotTargetingDrugsVSsimilarPerturbations(
predicted, compareCompounds, column="spearman_rank", labelBy="target")## Columns 'broad id' and 'pubchem_cid' were matched based on 0 common values; to manually select columns to compare, please set arguments starting with 'keyCol'## Warning: No shared levels found between `names(values)` of the manual scale and the
## data's colour values.
# Label by compound's mechanism of action (MOA)
plotTargetingDrugsVSsimilarPerturbations(
predicted, compareCompounds, column="spearman_rank", labelBy="moa")## Columns 'broad id' and 'pubchem_cid' were matched based on 0 common values; to manually select columns to compare, please set arguments starting with 'keyCol'## Warning: No shared levels found between `names(values)` of the manual scale and the
## data's colour values.
Molecular descriptor enrichment analysis
Next, from our candidate targeting drugs, we will analyse the enrichment of 2D and 3D molecular descriptors based on CMap and NCI60 compounds. This allows to check if targeting drugs are particularly enriched in specific drug descriptors and may be useful to think about the relevance of these descriptors for targeting a phenotype of interest.
descriptors <- loadDrugDescriptors("CMap", "2D")## compound_descriptors_CMap_2D.qs not found: downloading data...
drugSets <- prepareDrugSets(descriptors)
dsea <- analyseDrugSetEnrichment(drugSets, predicted)## Matching compounds with those available in drug sets...## Ordering results by column 'rankProduct_rank'; to manually select column to order by, please set argument 'col'## Columns 'broad id' and 'pert_id' were matched based on 283 common values; to manually select columns to compare, please set arguments starting with 'keyCol'## Performing enrichment analysis...
# Plot the 6 most significant drug set enrichment results
plotDrugSetEnrichment(drugSets, predicted,
selectedSets=head(dsea$descriptor, 6))## Matching compounds with those available in drug sets...## Columns 'broad id' and 'pert_id' were matched based on 283 common values; to manually select columns to compare, please set arguments starting with 'keyCol'## Plotting enrichment analysis...## $`Total Surface Area: [293, 307]`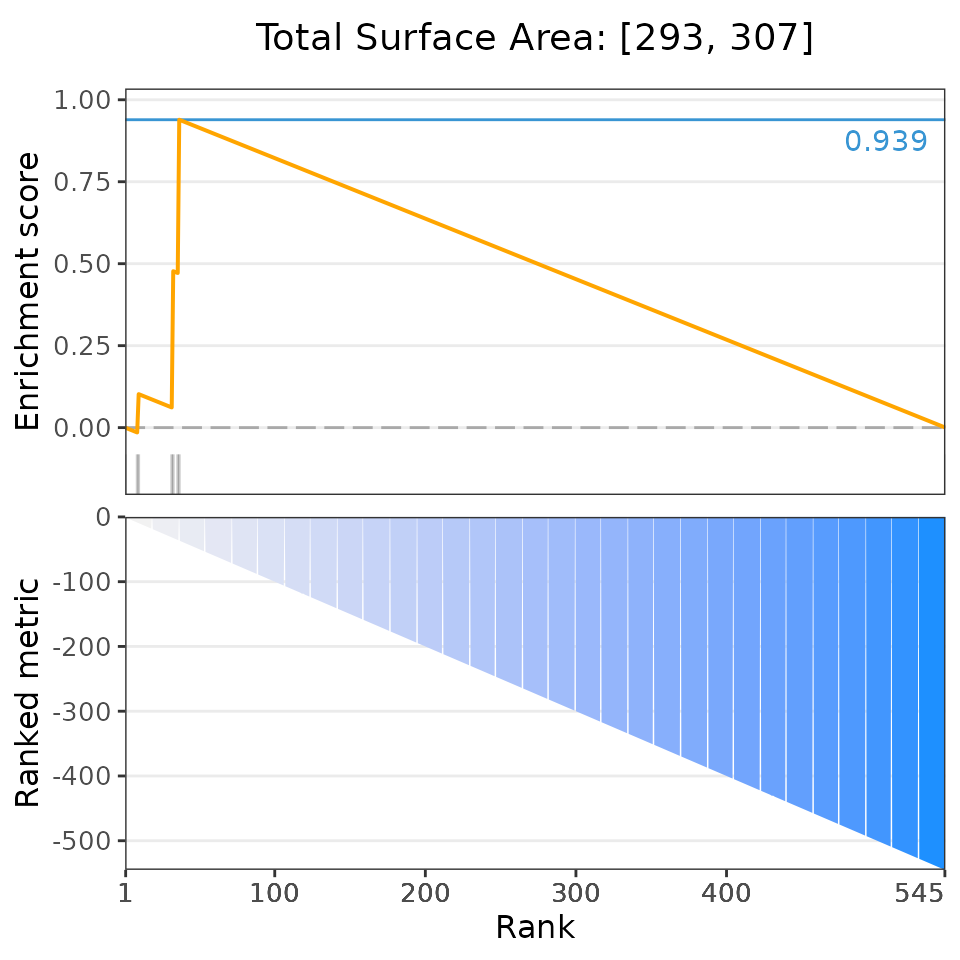
##
## $`Total.Molweight: [391, 411]`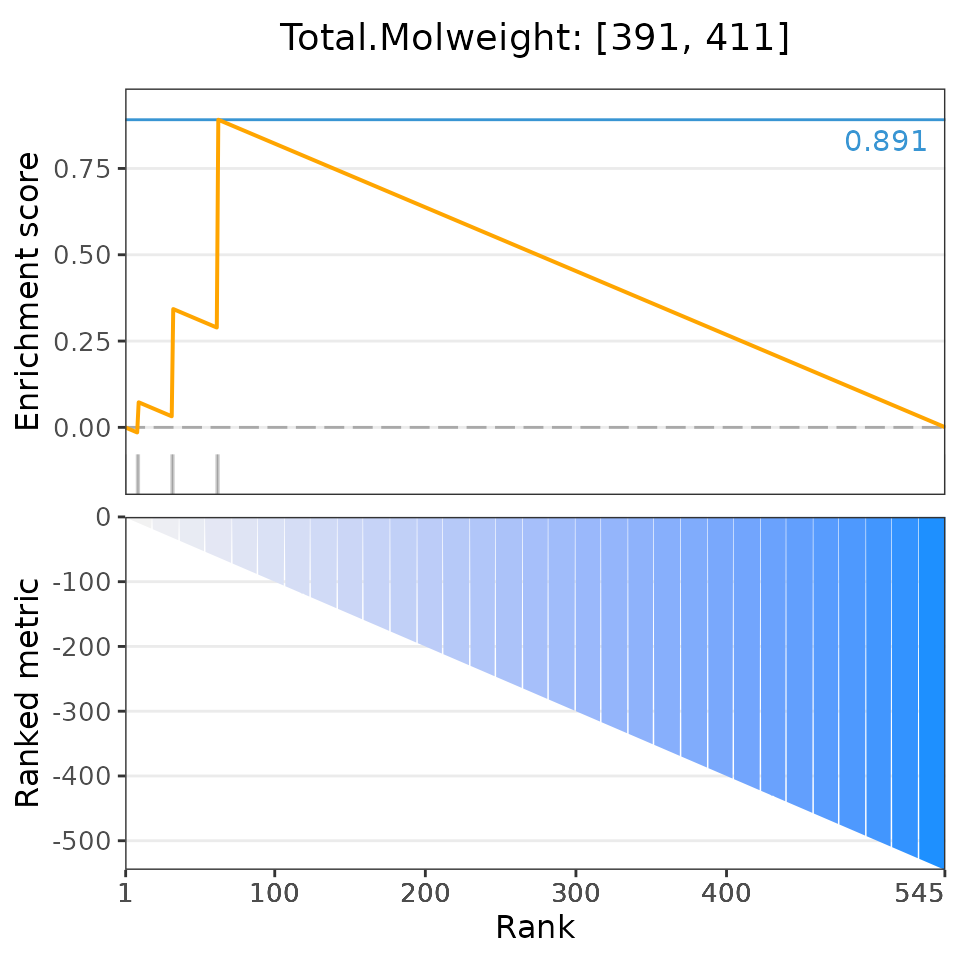
##
## $`Total Molweight: [391, 411]`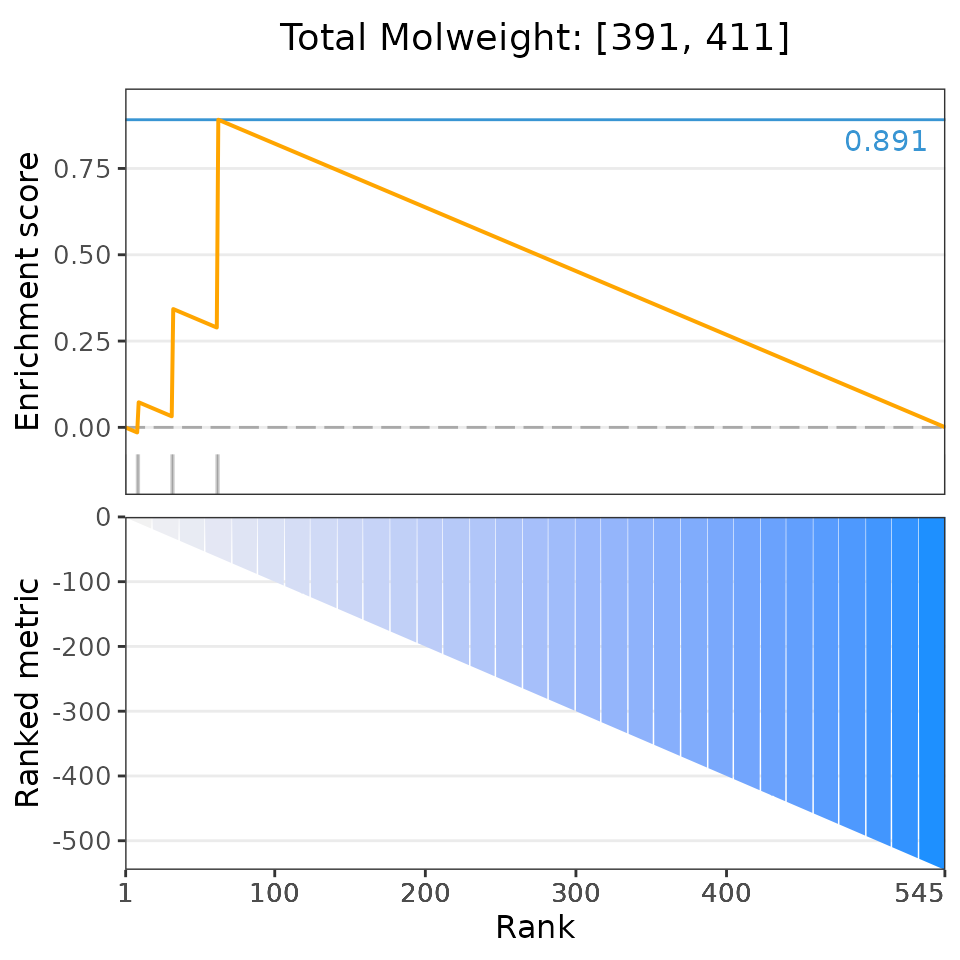
##
## $`Rotatable Bonds: 5`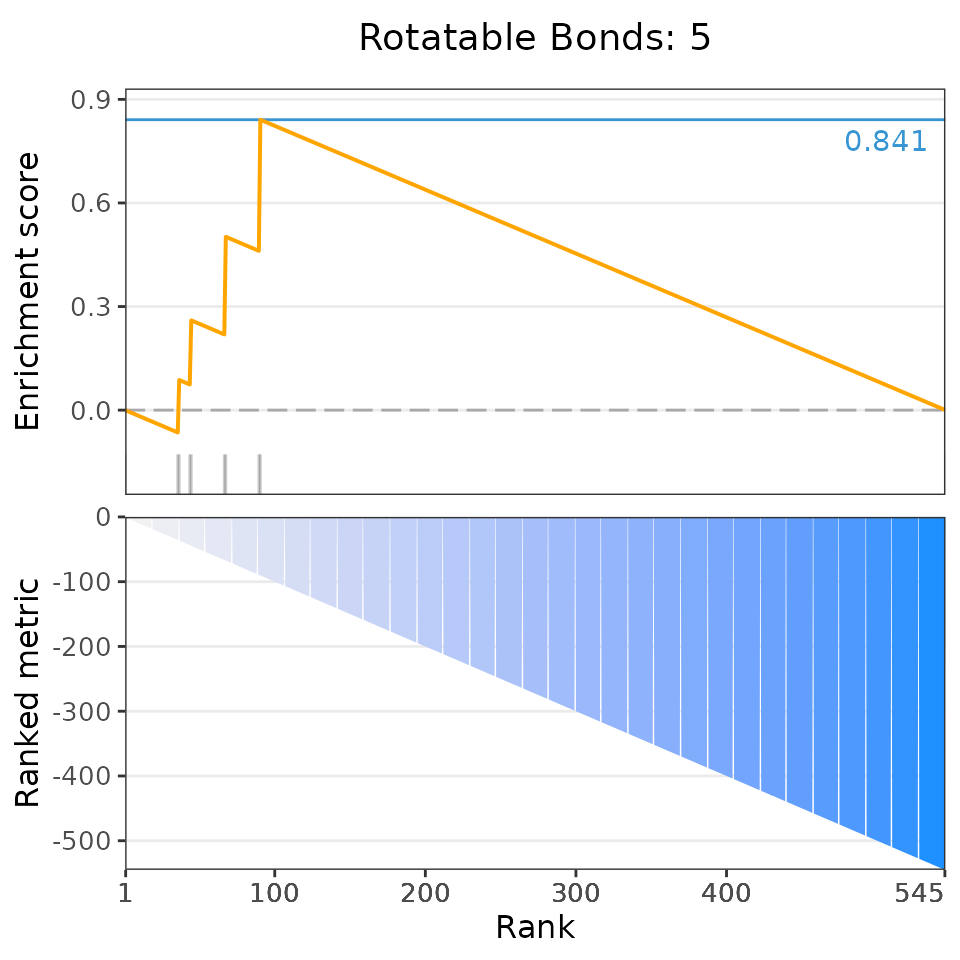
##
## $`Polar.Surface.Area: [138, 1153]`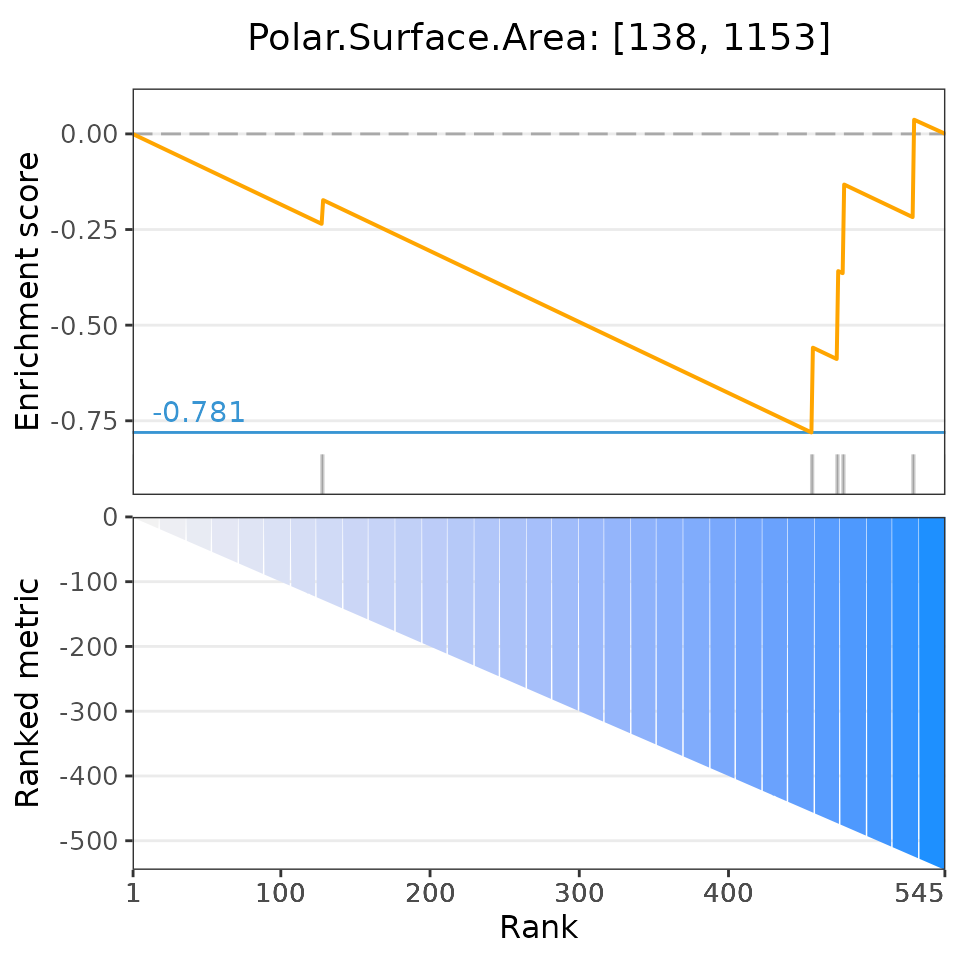
##
## $`cLogP: 0`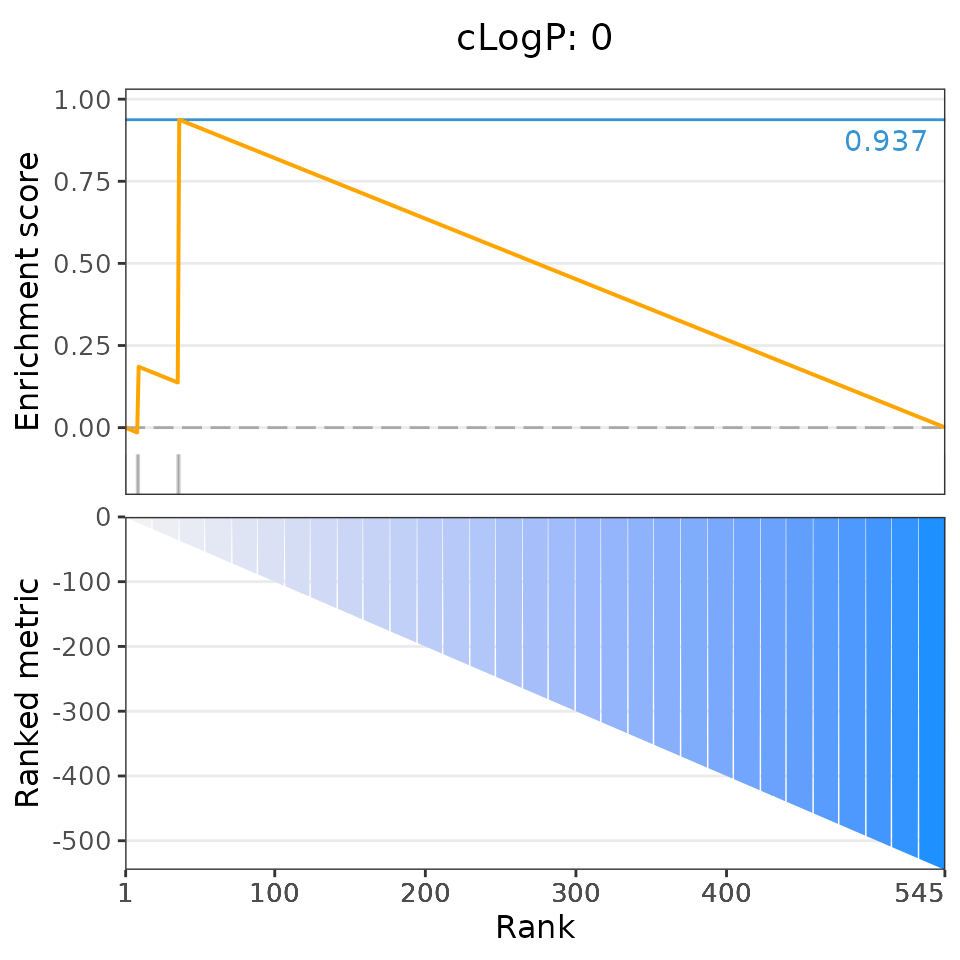
Contact information
All feedback on the program, documentation and associated material (including this tutorial) is welcome. Please send any suggestions and comments to:
Nuno Saraiva-Agostinho (nunoagostinho@medicina.ulisboa.pt)
Bernardo P. de Almeida (bernardo.almeida94@gmail.com)
Disease Transcriptomics Lab, Instituto de Medicina Molecular (Portugal)